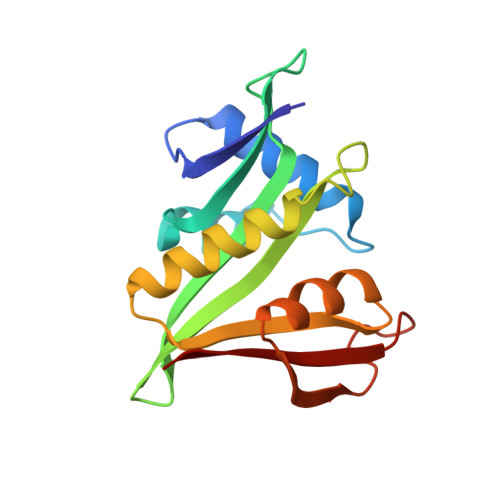
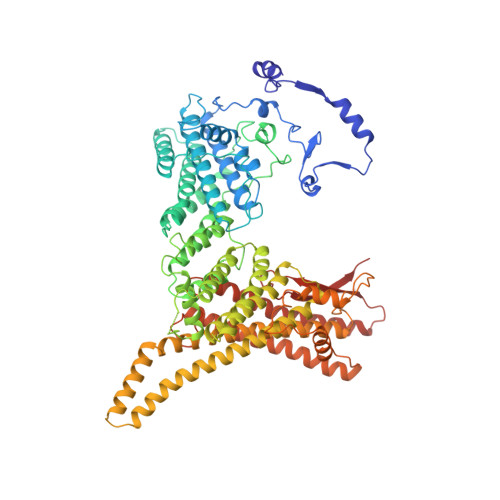
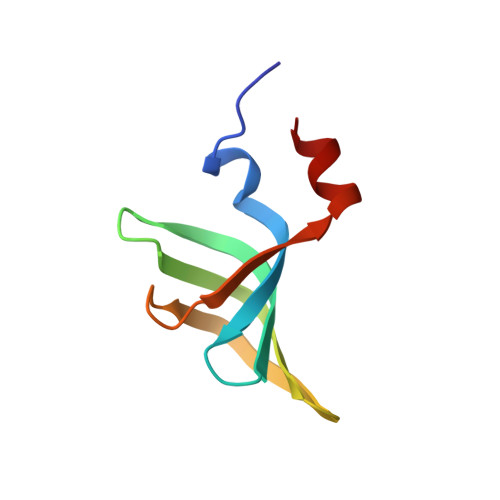
Divergent architecture of the heterotrimeric NatC complex explains N-terminal acetylation of cognate substrates.
Grunwald, S., Hopf, L.V.M., Bock-Bierbaum, T., Lally, C.C.M., Spahn, C.M.T., Daumke, O.(2020) Nat Commun 11: 5506-5506
- PubMed: 33139728 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-020-19321-8
- Primary Citation Related Structures:
6YGA, 6YGB, 6YGC, 6YGD - PubMed Abstract:
The heterotrimeric NatC complex, comprising the catalytic Naa30 and the two auxiliary subunits Naa35 and Naa38, co-translationally acetylates the N-termini of numerous eukaryotic target proteins. Despite its unique subunit composition, its essential role for many aspects of cellular function and its suggested involvement in disease, structure and mechanism of NatC have remained unknown. Here, we present the crystal structure of the Saccharomyces cerevisiae NatC complex, which exhibits a strikingly different architecture compared to previously described N-terminal acetyltransferase (NAT) complexes. Cofactor and ligand-bound structures reveal how the first four amino acids of cognate substrates are recognized at the Naa30-Naa35 interface. A sequence-specific, ligand-induced conformational change in Naa30 enables efficient acetylation. Based on detailed structure-function studies, we suggest a catalytic mechanism and identify a ribosome-binding patch in an elongated tip region of NatC. Our study reveals how NAT machineries have divergently evolved to N-terminally acetylate specific subsets of target proteins.
- Department of Crystallography, Max Delbrück Center for Molecular Medicine, 13125, Berlin, Germany.
Organizational Affiliation: