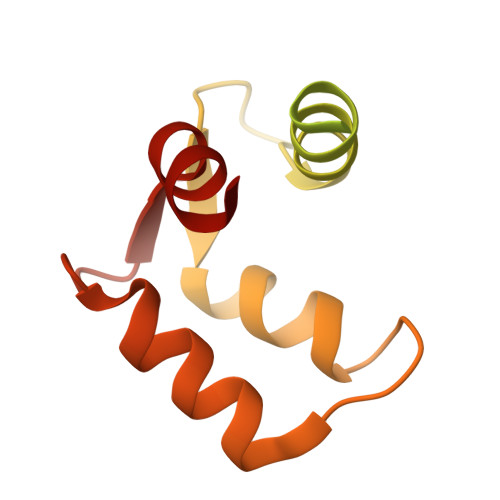
Activation mechanism of a human SK-calmodulin channel complex elucidated by cryo-EM structures.
Lee, C.H., MacKinnon, R.(2018) Science 360: 508-513
- PubMed: 29724949
- DOI: https://doi.org/10.1126/science.aas9466
- Primary Citation of Related Structures:
6CNM, 6CNN, 6CNO - PubMed Abstract:
Small-conductance Ca 2+ -activated K + (SK) channels mediate neuron excitability and are associated with synaptic transmission and plasticity. They also regulate immune responses and the size of blood cells. Activation of SK channels requires calmodulin (CaM), but how CaM binds and opens SK channels has been unclear. Here we report cryo-electron microscopy (cryo-EM) structures of a human SK4-CaM channel complex in closed and activated states at 3.4- and 3.5-angstrom resolution, respectively. Four CaM molecules bind to one channel tetramer. Each lobe of CaM serves a distinct function: The C-lobe binds to the channel constitutively, whereas the N-lobe interacts with the S4-S5 linker in a Ca 2+ -dependent manner. The S4-S5 linker, which contains two distinct helices, undergoes conformational changes upon CaM binding to open the channel pore. These structures reveal the gating mechanism of SK channels and provide a basis for understanding SK channel pharmacology.
- Laboratory of Molecular Neurobiology and Biophysics, The Rockefeller University, Howard Hughes Medical Institute, 1230 York Avenue, New York, NY 10065, USA.
Organizational Affiliation: