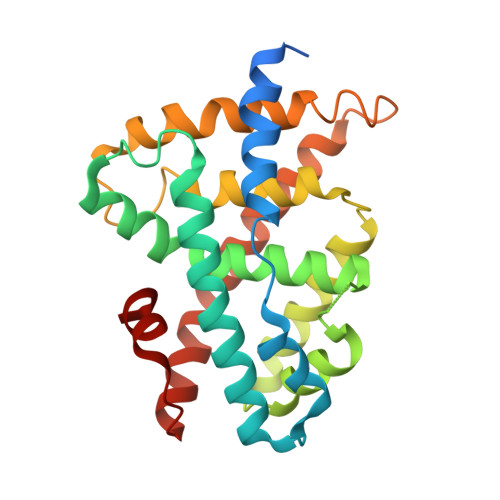
Structural basis of tropifexor as a potent and selective agonist of farnesoid X receptor.
Jiang, L., Xiao, D., Li, Y., Dai, S., Qu, L., Chen, X., Guo, M., Wei, H., Chen, Y.(2021) Biochem Biophys Res Commun 534: 1047-1052
- PubMed: 33121679 Search on PubMed
- DOI: https://doi.org/10.1016/j.bbrc.2020.10.039
- Primary Citation Related Structures:
7D42 - PubMed Abstract:
Farnesoid X receptor (FXR) is considered as a potential target for the treatment of several liver disorders such as primary biliary cholangitis (PBC) and primary sclerosing cholangitis (PSC). Tropifexor is a highly potent and non-steroidal FXR agonist that has progressed into phase II clinical trials in patients with PBC. The clinical trials demonstrate that tropifexor improved serum markers of patients with liver diseases and lower side effect such as pruritus that might be implicated with TGR5 activation. However, the molecular mechanism of the potency and selectivity of tropifexor remains unclear. In this study, the binding affinity of FXR and tropifexor is measured by isothermal titration calorimetry (ITC) assays. The crystal structure of the FXR/tropifexor complex is determined at 2.7 Å resolution to explain the molecular mechanism of tropifexor bound to FXR-LBD. Structural comparison with other FXR/agonists structures reveals the conformational change in the FXR/tropifexor structure. Moreover the structural superposition of TGR5/tropifexor indicates that the steric hindrance between tropifexor and TGR5 might be a possible explanation for the impotency arises of tropifexor to TGR5. Overall, our analyses might provide an insight into the molecular mechanism of tropifexor binding to FXR-LBD and account for the high selectivity of tropifexor for FXR versus TGR5.
- Department of Pathology, NHC Key Laboratory of Cancer Proteomics, Laboratory of Structural Biology, National Clinical Research Center for Geriatric Disorders, Xiangya Hospital, Central South University, Changsha, Hunan, 410008, China.
Organizational Affiliation: