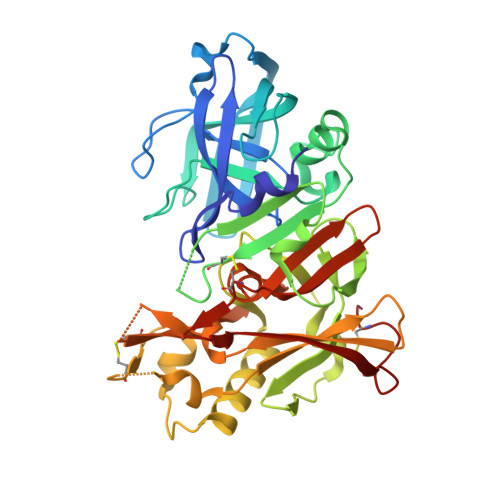
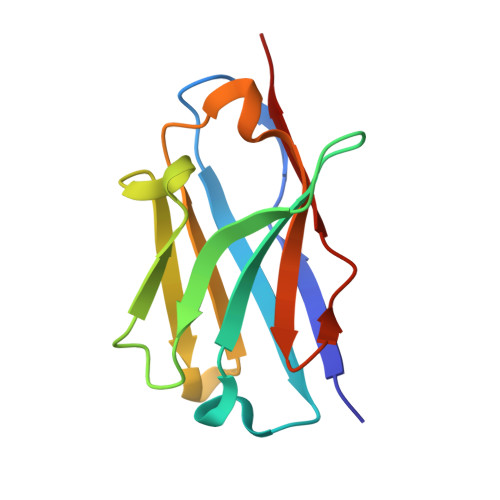
Structure-Based Approaches to Improving Selectivity through Utilizing Explicit Water Molecules: Discovery of Selective beta-Secretase (BACE1) Inhibitors over BACE2.
Fujimoto, K., Yoshida, S., Tadano, G., Asada, N., Fuchino, K., Suzuki, S., Matsuoka, E., Yamamoto, T., Yamamoto, S., Ando, S., Kanegawa, N., Tonomura, Y., Ito, H., Moechars, D., Rombouts, F.J.R., Gijsen, H.J.M., Kusakabe, K.I.(2021) J Med Chem 64: 3075-3085
- PubMed: 33719429
- DOI: https://doi.org/10.1021/acs.jmedchem.0c01858
- Primary Citation of Related Structures:
7D2V, 7D2X, 7D5A, 7D5B, 7D5U - PubMed Abstract:
BACE1 is an attractive target for disease-modifying treatment of Alzheimer's disease. BACE2, having high homology around the catalytic site, poses a critical challenge to identifying selective BACE1 inhibitors. Recent evidence indicated that BACE2 has various roles in peripheral tissues and the brain, and therefore, the chronic use of nonselective inhibitors may cause side effects derived from BACE2 inhibition. Crystallographic analysis of the nonselective inhibitor verubecestat identified explicit water molecules with different levels of free energy in the S2' pocket. Structure-based design targeting them enabled the identification of propynyl oxazine 3 with improved selectivity. Further optimization efforts led to the discovery of compound 6 with high selectivity. The cocrystal structures of 7 , a close analogue of 6 , bound to BACE1 and BACE2 confirmed that one of the explicit water molecules is displaced by the propynyl group, suggesting that the difference in the relative water displacement cost may contribute to the improved selectivity.
- Laboratory for Medicinal Chemistry Research, Shionogi Pharmaceutical Research Center, 1-1 Futaba-cho 3-chome, Toyonaka, Osaka 561-0825, Japan.
Organizational Affiliation: