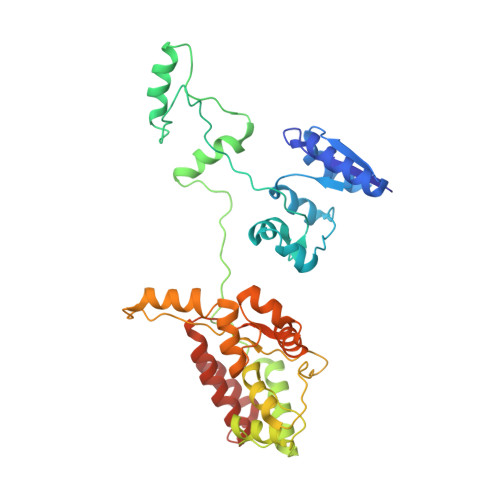
Crystal structure of the domain-swapped dimeric maltodextrin-binding protein MalE from Salmonella enterica.
Wang, L., Bu, T., Bai, X., He, S., Zhang, J., Jin, L., Liu, B., Dong, Y., Ha, N.C., Quan, C., Nam, K.H., Xu, Y.(2022) Acta Crystallogr D Struct Biol 78: 613-622
- PubMed: 35503209 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798322003114
- Primary Citation Related Structures:
7FFT, 7FFW - PubMed Abstract:
MalE is a maltose/maltodextrin-binding protein (MBP) that plays a critical role in most bacterial maltose/maltodextrin-transport systems. Previously reported wild-type MBPs are monomers comprising an N-terminal domain (NTD) and a C-terminal domain (CTD), and maltose-like molecules are recognized between the NTD and CTD and transported to the cell system. Because MBP does not undergo artificial dimerization, it is widely used as a tag for protein expression and purification. Here, the crystal structure of a domain-swapped dimeric MalE from Salmonella enterica (named SeMalE) in complex with maltopentaose is reported for the first time, and its structure is distinct from typical monomeric MalE family members. In the domain-swapped dimer, SeMalE comprises two subdomains: the NTD and CTD. The NTD and CTD of one molecule of SeMalE interact with the CTD and NTD of the partner molecule, respectively. The domain-swapped dimeric conformation was stabilized by interactions between the NTDs, CTDs and linkers from two SeMalE molecules. Additionally, a maltopentaose molecule was found to be located at the interface between the NTD and CTD of different SeMalE molecules. These results provide new insights that will improve the understanding of maltodextrin-binding MalE proteins.
- Department of Bioengineering, College of Life Science, Dalian Minzu University, No. 18 Liaohe West Road, Dalian, 116600 Liaoning, People's Republic of China.
Organizational Affiliation: