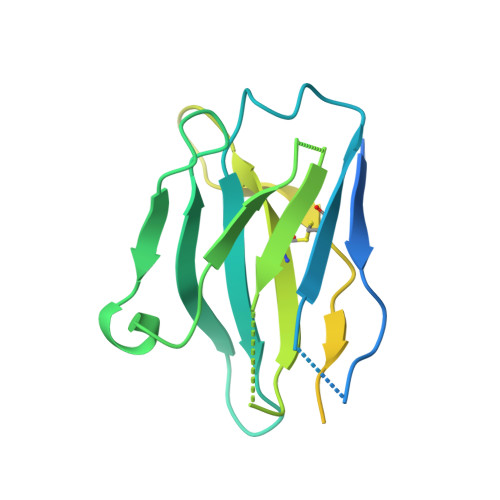
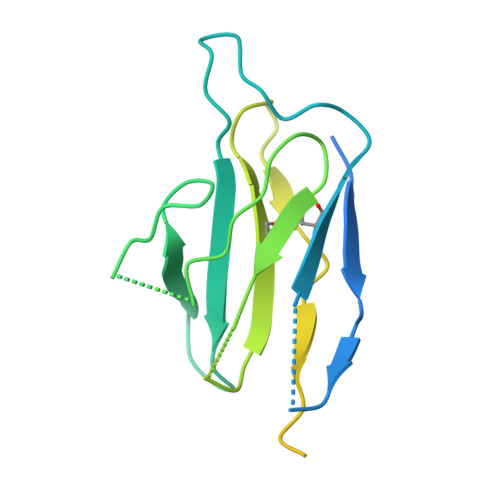
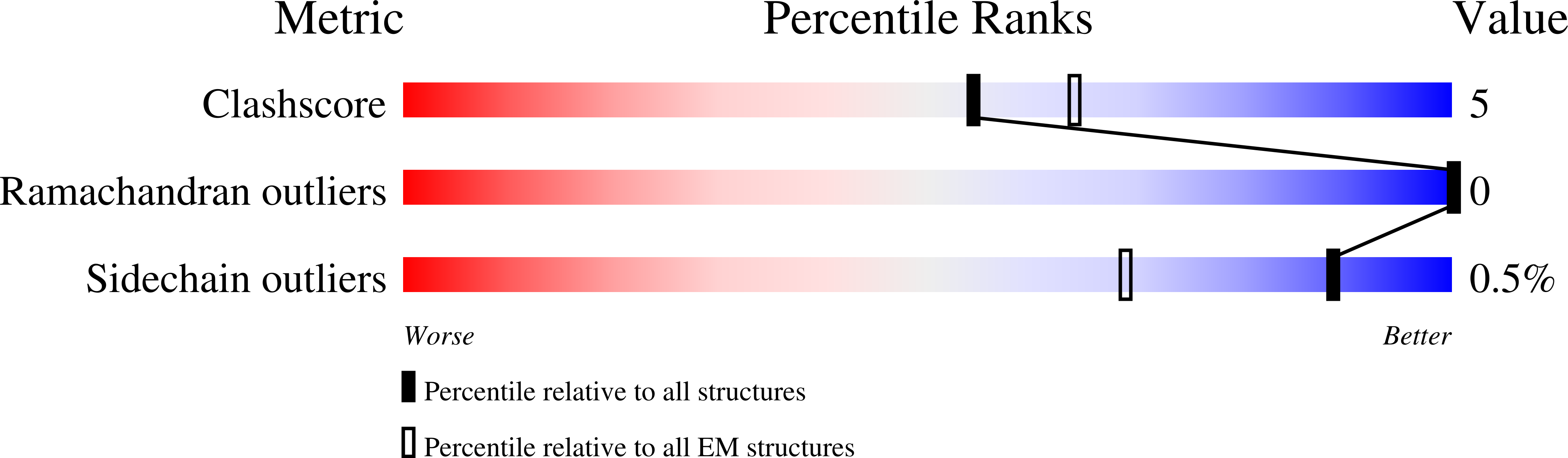Substrate and product complexes reveal mechanisms of Hedgehog acylation by HHAT.
Jiang, Y., Benz, T.L., Long, S.B.(2021) Science 372: 1215-1219
- PubMed: 34112694
- DOI: https://doi.org/10.1126/science.abg4998
- Primary Citation of Related Structures:
7MHY, 7MHZ - PubMed Abstract:
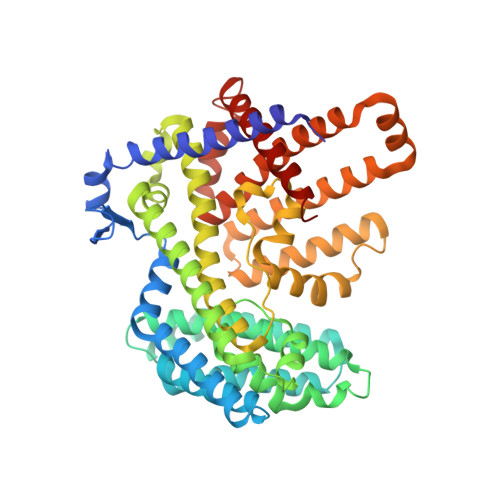
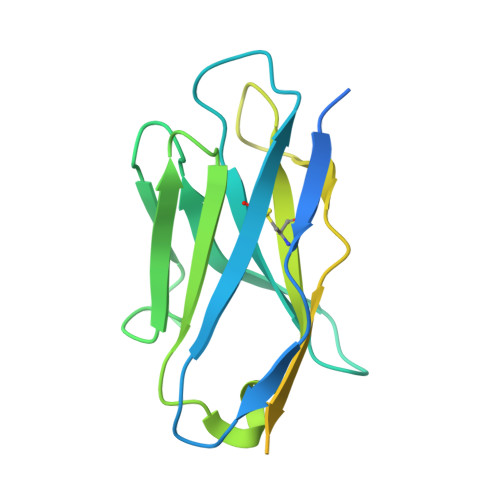
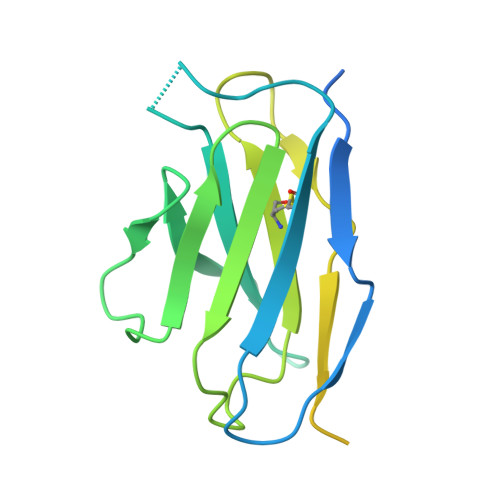
Hedgehog proteins govern crucial developmental steps in animals and drive certain human cancers. Before they can function as signaling molecules, Hedgehog precursor proteins must undergo amino-terminal palmitoylation by Hedgehog acyltransferase (HHAT). We present cryo-electron microscopy structures of human HHAT in complex with its palmitoyl-coenzyme A substrate and of a product complex with a palmitoylated Hedgehog peptide at resolutions of 2.7 and 3.2 angstroms, respectively. The structures reveal how HHAT overcomes the challenges of bringing together substrates that have different physiochemical properties from opposite sides of the endoplasmic reticulum membrane within a membrane-embedded active site for catalysis. These principles are relevant to related enzymes that catalyze the acylation of Wnt and of the appetite-stimulating hormone ghrelin. The structural and mechanistic insights may advance the development of inhibitors for cancer.
Organizational Affiliation:
Structural Biology Program, Memorial Sloan Kettering Cancer Center, New York, NY 10065, USA.