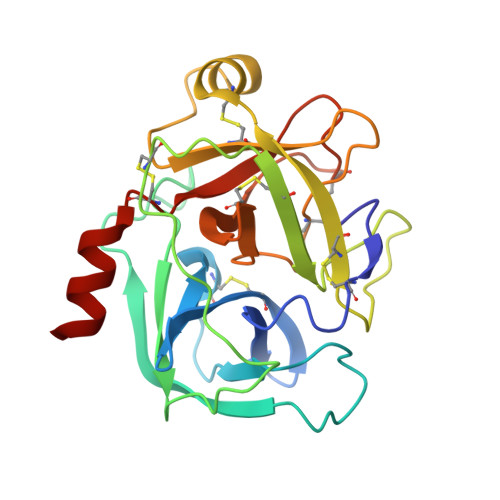
Scalable synthesis and structural characterization of reversible KLK6 inhibitors.
Baumann, A., Isak, D., Lohbeck, J., Jagtap, P.K.A., Hennig, J., Miller, A.K.(2022) RSC Adv 12: 26989-26993
- PubMed: 36320846
- DOI: https://doi.org/10.1039/d2ra04670a
- Primary Citation of Related Structures:
7QHZ, 7QI0 - PubMed Abstract:
Scalable asymmetric syntheses of two kallikrein-related protease 6 (KLK6) inhibitors are reported. The inhibitors are assembled by linking enantiomerically enriched fragments via amide bond formation, followed by conversion of a cyano group to an amidine. One fragment, an amine, was prepared using the Ellman auxiliary, and a lack of clarity in the literature regarding the stereochemical outcome of this reaction was solved via X-ray crystallographic analysis of two derivatives. Complexes of the inhibitors bound to human KLK6 were solved by X-ray crystallography, revealing the binding poses.
Organizational Affiliation:
Cancer Drug Development Group, German Cancer Research Center (DKFZ) Im Neuenheimer Feld 280 69120 Heidelberg Germany aubry.miller@dkfz.de.