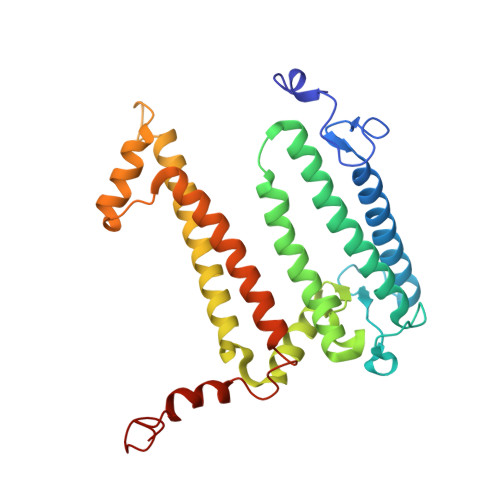
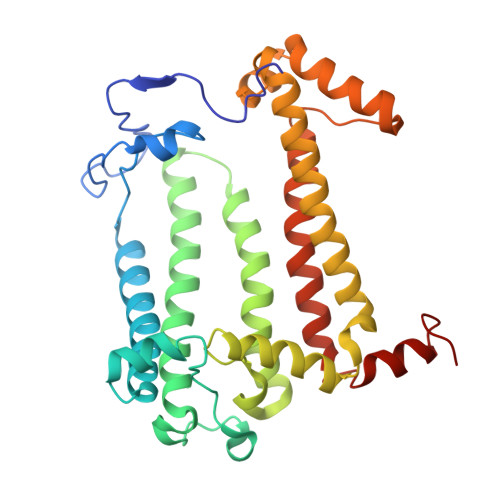
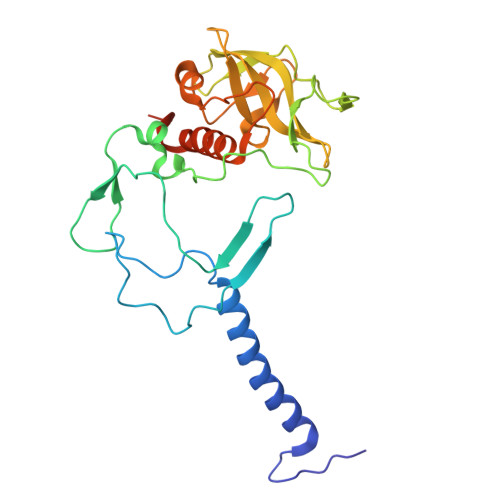
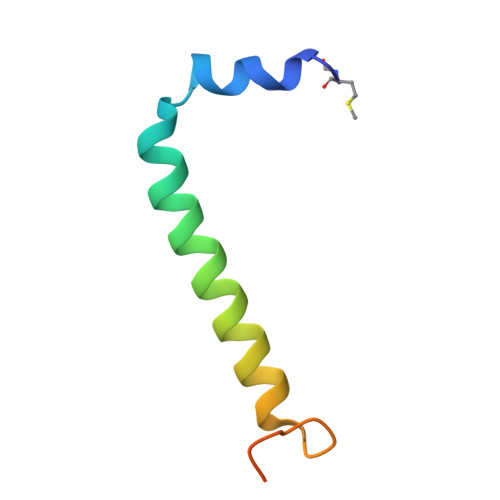
Asymmetric structure of the native Rhodobacter sphaeroides dimeric LH1-RC complex.
Tani, K., Kanno, R., Kikuchi, R., Kawamura, S., Nagashima, K.V.P., Hall, M., Takahashi, A., Yu, L.J., Kimura, Y., Madigan, M.T., Mizoguchi, A., Humbel, B.M., Wang-Otomo, Z.Y.(2022) Nat Commun 13: 1904-1904
- PubMed: 35393413 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-022-29453-8
- Primary Citation Related Structures:
7VY2, 7VY3 - PubMed Abstract:
Rhodobacter sphaeroides is a model organism in bacterial photosynthesis, and its light-harvesting-reaction center (LH1-RC) complex contains both dimeric and monomeric forms. Here we present cryo-EM structures of the native LH1-RC dimer and an LH1-RC monomer lacking protein-U (ΔU). The native dimer reveals several asymmetric features including the arrangement of its two monomeric components, the structural integrity of protein-U, the overall organization of LH1, and rigidities of the proteins and pigments. PufX plays a critical role in connecting the two monomers in a dimer, with one PufX interacting at its N-terminus with another PufX and an LH1 β-polypeptide in the other monomer. One protein-U was only partially resolved in the dimeric structure, signaling different degrees of disorder in the two monomers. The ΔU LH1-RC monomer was half-moon-shaped and contained 11 α- and 10 β-polypeptides, indicating a critical role for protein-U in controlling the number of αβ-subunits required for dimer assembly and stabilization. These features are discussed in relation to membrane topology and an assembly model proposed for the native dimeric complex.
- Graduate School of Medicine, Mie University, Tsu, 514-8507, Japan. ktani@doc.medic.mie-u.ac.jp.
Organizational Affiliation: