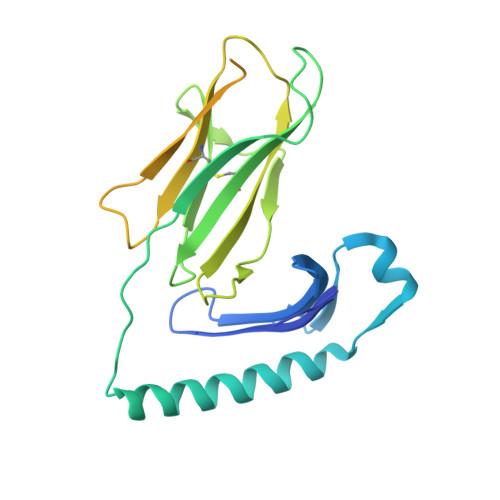
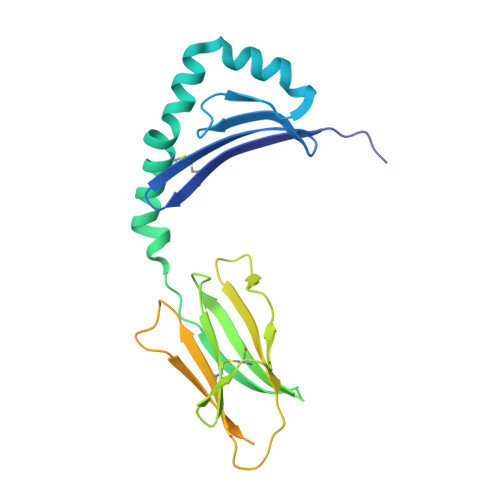
Machine learning predictions of MHC-II specificities reveal alternative binding mode of class II epitopes.
Racle, J., Guillaume, P., Schmidt, J., Michaux, J., Larabi, A., Lau, K., Perez, M.A.S., Croce, G., Genolet, R., Coukos, G., Zoete, V., Pojer, F., Bassani-Sternberg, M., Harari, A., Gfeller, D.(2023) Immunity 56: 1359-1375.e13
- PubMed: 37023751
- DOI: https://doi.org/10.1016/j.immuni.2023.03.009
- Primary Citation Related Structures:
7ZAK, 7ZFR - PubMed Abstract:
CD4 + T cells orchestrate the adaptive immune response against pathogens and cancer by recognizing epitopes presented on class II major histocompatibility complex (MHC-II) molecules. The high polymorphism of MHC-II genes represents an important hurdle toward accurate prediction and identification of CD4 + T cell epitopes. Here we collected and curated a dataset of 627,013 unique MHC-II ligands identified by mass spectrometry. This enabled us to precisely determine the binding motifs of 88 MHC-II alleles across humans, mice, cattle, and chickens. Analysis of these binding specificities combined with X-ray crystallography refined our understanding of the molecular determinants of MHC-II motifs and revealed a widespread reverse-binding mode in HLA-DP ligands. We then developed a machine-learning framework to accurately predict binding specificities and ligands of any MHC-II allele. This tool improves and expands predictions of CD4 + T cell epitopes and enables us to discover viral and bacterial epitopes following the aforementioned reverse-binding mode.
- Department of Oncology UNIL CHUV, Ludwig Institute for Cancer Research, University of Lausanne, Lausanne, Switzerland; Swiss Institute of Bioinformatics (SIB), Lausanne, Switzerland; Agora Cancer Research Centre, Lausanne, Switzerland; Swiss Cancer Center Leman (SCCL), Lausanne, Switzerland. Electronic address: julien.racle@unil.ch.
Organizational Affiliation: