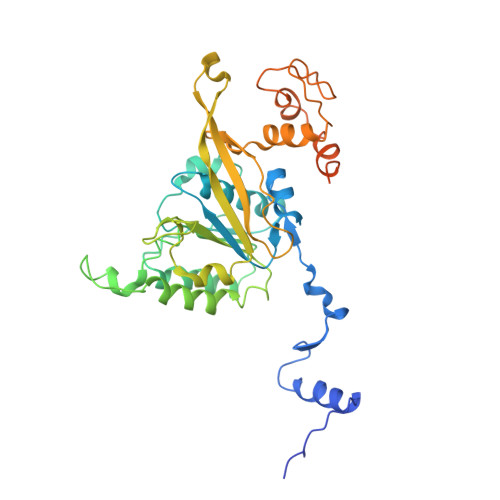
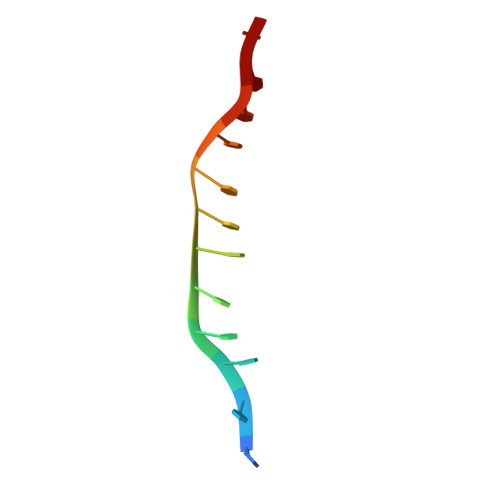
Assembly mechanism and cryoEM structure of RecA recombination nucleofilaments from Streptococcus pneumoniae.
Hertzog, M., Perry, T.N., Dupaigne, P., Serres, S., Morales, V., Soulet, A.L., Bell, J.C., Margeat, E., Kowalczykowski, S.C., Le Cam, E., Fronzes, R., Polard, P.(2023) Nucleic Acids Res 51: 2800-2817
- PubMed: 36806960 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1093/nar/gkad080
- Primary Citation Related Structures:
8AMD, 8AMF - PubMed Abstract:
RecA-mediated homologous recombination (HR) is a key mechanism for genome maintenance and plasticity in bacteria. It proceeds through RecA assembly into a dynamic filament on ssDNA, the presynaptic filament, which mediates DNA homology search and ordered DNA strand exchange. Here, we combined structural, single molecule and biochemical approaches to characterize the ATP-dependent assembly mechanism of the presynaptic filament of RecA from Streptococcus pneumoniae (SpRecA), in comparison to the Escherichia coli RecA (EcRecA) paradigm. EcRecA polymerization on ssDNA is assisted by the Single-Stranded DNA Binding (SSB) protein, which unwinds ssDNA secondary structures that block EcRecA nucleofilament growth. We report by direct microscopic analysis of SpRecA filamentation on ssDNA that neither of the two paralogous pneumococcal SSBs could assist the extension of SpRecA nucleopolymers. Instead, we found that the conserved RadA helicase promotes SpRecA nucleofilamentation in an ATP-dependent manner. This allowed us to solve the atomic structure of such a long native SpRecA nucleopolymer by cryoEM stabilized with ATPγS. It was found to be equivalent to the crystal structure of the EcRecA filament with a marked difference in how RecA mediates nucleotide orientation in the stretched ssDNA. Then, our results show that SpRecA and EcRecA HR activities are different, in correlation with their distinct ATP-dependent ssDNA binding modes.
- Laboratoire de Microbiologie et de Génétique Moléculaire (UMR 5100). Centre de Biologie Intégrative; 169, avenue Marianne Grunberg-Manago; CNRS - Université Paul Sabatier - Bât 4R4, 118, route de Narbonne; 31062 Toulouse cedex 09, France.
Organizational Affiliation: