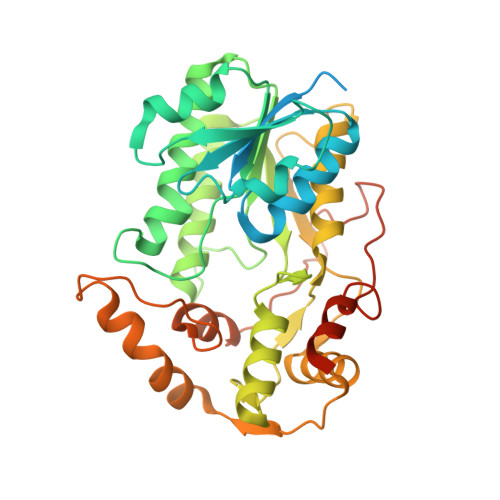
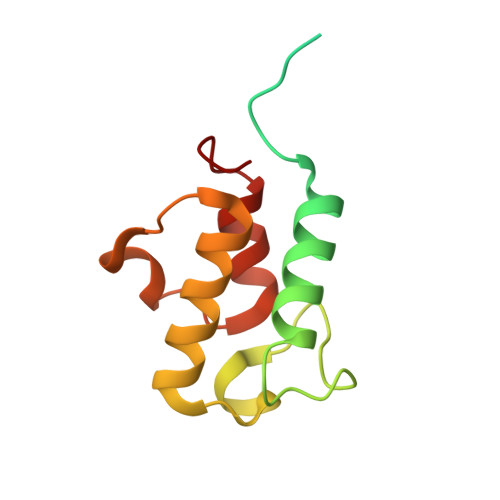
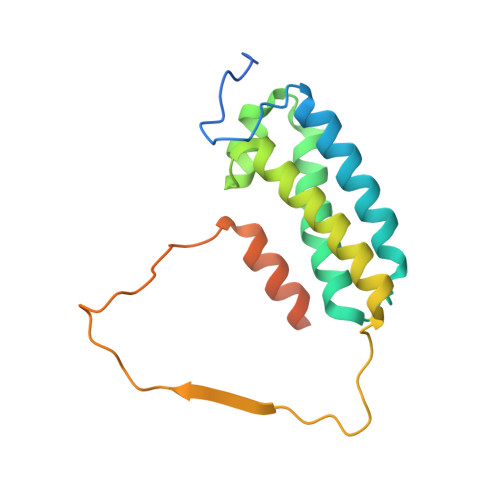
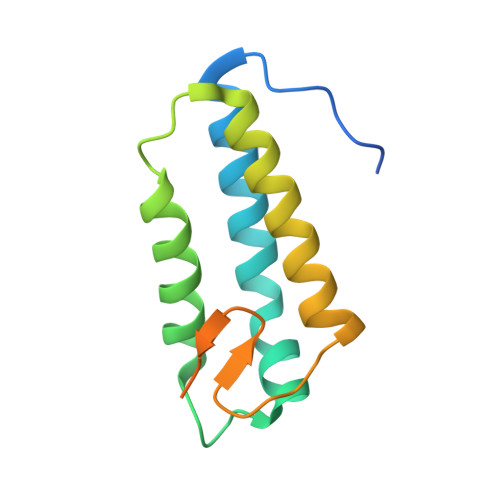
Cryo-EM structure of the respiratory I + III 2 supercomplex from Arabidopsis thaliana at 2 angstrom resolution.
Klusch, N., Dreimann, M., Senkler, J., Rugen, N., Kuhlbrandt, W., Braun, H.P.(2023) Nat Plants 9: 142-156
- PubMed: 36585502
- DOI: https://doi.org/10.1038/s41477-022-01308-6
- Primary Citation Related Structures:
8BED, 8BEE, 8BEF, 8BEH, 8BEL, 8BEP, 8BPX, 8BQ5, 8BQ6 - PubMed Abstract:
Protein complexes of the mitochondrial respiratory chain assemble into respiratory supercomplexes. Here we present the high-resolution electron cryo-microscopy structure of the Arabidopsis respiratory supercomplex consisting of complex I and a complex III dimer, with a total of 68 protein subunits and numerous bound cofactors. A complex I-ferredoxin, subunit B14.7 and P9, a newly defined subunit of plant complex I, mediate supercomplex formation. The component complexes stabilize one another, enabling new detailed insights into their structure. We describe (1) an interrupted aqueous passage for proton translocation in the membrane arm of complex I; (2) a new coenzyme A within the carbonic anhydrase module of plant complex I defining a second catalytic centre; and (3) the water structure at the proton exit pathway of complex III 2 with a co-purified ubiquinone in the Q O site. We propose that the main role of the plant supercomplex is to stabilize its components in the membrane.
- Department of Structural Biology, Max-Planck-Institute of Biophysics, Frankfurt, Germany. Niklas.Klusch@biophys.mpg.de.
Organizational Affiliation: