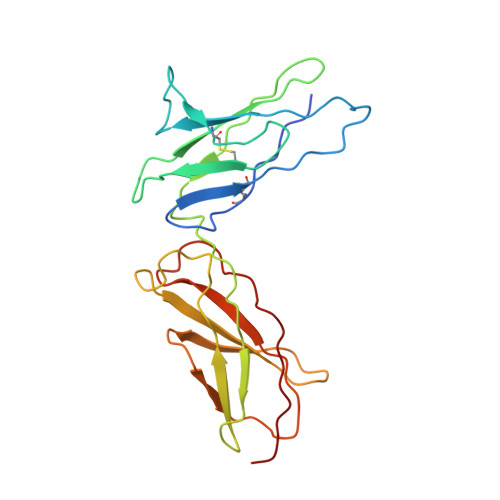
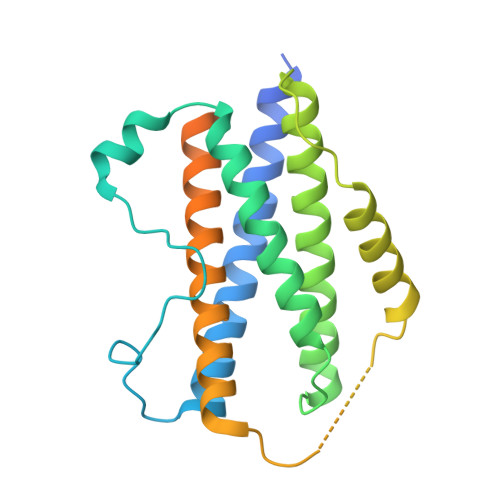
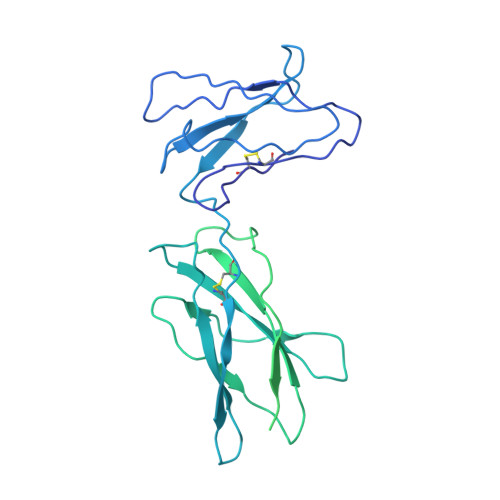
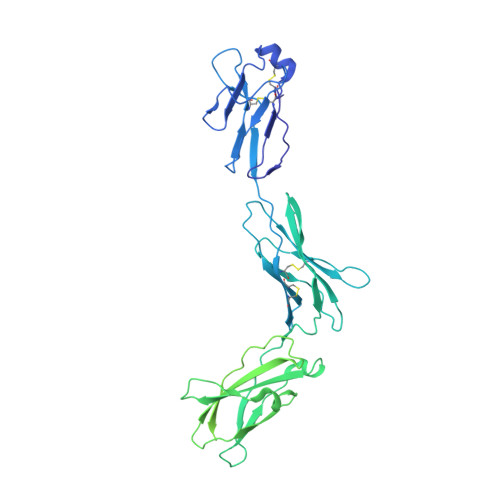
Structural insights into the assembly of gp130 family cytokine signaling complexes.
Zhou, Y., Stevis, P.E., Cao, J., Saotome, K., Wu, J., Glatman Zaretsky, A., Haxhinasto, S., Yancopoulos, G.D., Murphy, A.J., Sleeman, M.W., Olson, W.C., Franklin, M.C.(2023) Sci Adv 9: eade4395-eade4395
- PubMed: 36930708 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.ade4395
- Primary Citation Related Structures:
8D6A, 8D74, 8D7E, 8D7H, 8D7R, 8D82, 8D85 - PubMed Abstract:
The interleukin-6 (IL-6) family cytokines signal through gp130 receptor homodimerization or heterodimerization with a second signaling receptor and play crucial roles in various cellular processes. We determined cryo-electron microscopy structures of five signaling complexes of this family, containing full receptor ectodomains bound to their respective ligands ciliary neurotrophic factor, cardiotrophin-like cytokine factor 1 (CLCF1), leukemia inhibitory factor, IL-27, and IL-6. Our structures collectively reveal similarities and differences in the assembly of these complexes. The acute bends at both signaling receptors in all complexes bring the membrane-proximal domains to a ~30 angstrom range but with distinct distances and orientations. We also reveal how CLCF1 engages its secretion chaperone cytokine receptor-like factor 1. Our data provide valuable insights for therapeutically targeting gp130-mediated signaling.
- Regeneron Pharmaceuticals Inc., Tarrytown, NY 10591, USA.
Organizational Affiliation: