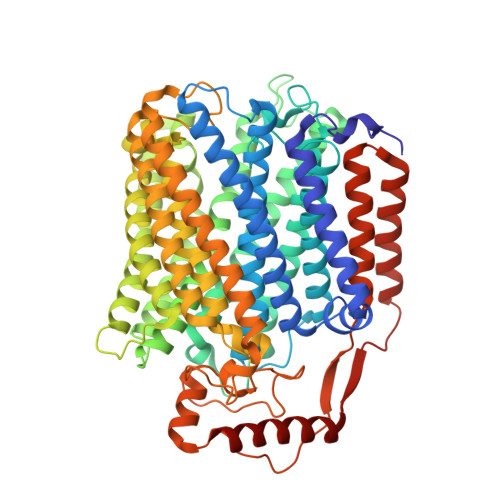
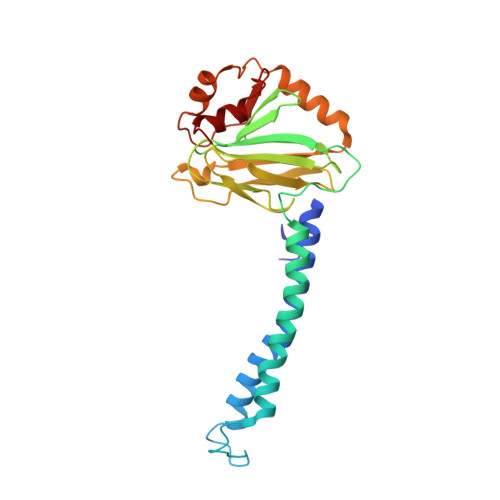
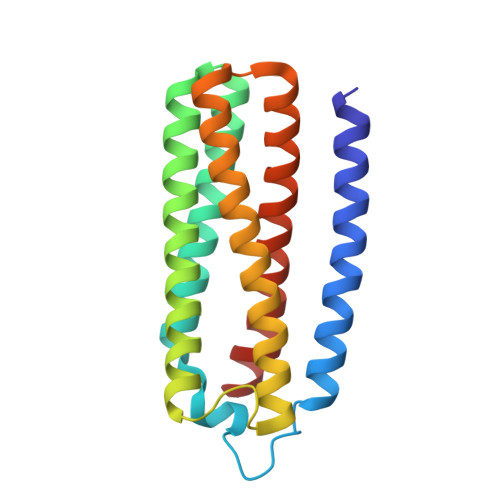
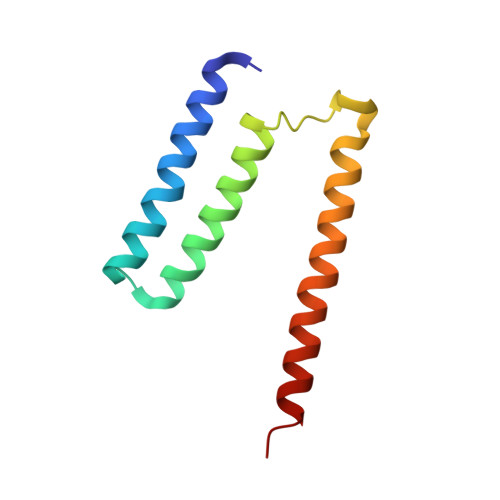
Monomer and dimer structures of cytochrome bo 3 ubiquinol oxidase from Escherichia coli.
Guo, Y., Karimullina, E., Emde, T., Otwinowski, Z., Borek, D., Savchenko, A.(2023) Protein Sci 32: e4616-e4616
- PubMed: 36880269 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.4616
- Primary Citation Related Structures:
8F68, 8F6C - PubMed Abstract:
The Escherichia coli cytochrome bo 3 ubiquinol oxidase is a four-subunit heme-copper oxidase that serves as a proton pump in the E. coli aerobic respiratory chain. Despite many mechanistic studies, it is unclear whether this ubiquinol oxidase functions as a monomer, or as a dimer in a manner similar to its eukaryotic counterparts-the mitochondrial electron transport complexes. In this study, we determined the monomeric and dimeric structures of the E. coli cytochrome bo 3 ubiquinol oxidase reconstituted in amphipol by cryogenic electron microscopy single particle reconstruction (cryo-EM SPR) to a resolution of 3.15 and 3.46 Å, respectively. We have discovered that the protein can form a dimer with C2 symmetry, with the dimerization interface maintained by interactions between the subunit II of one monomer and the subunit IV of the other monomer. Moreover, the dimerization does not induce significant structural changes in the monomers, except the movement of a loop in subunit IV (residues 67-74).
- Department of Biophysics, The University of Texas Southwestern Medical Center, Dallas, Texas, USA.
Organizational Affiliation: