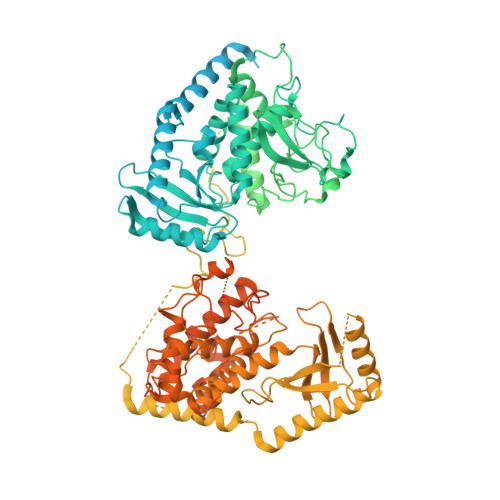
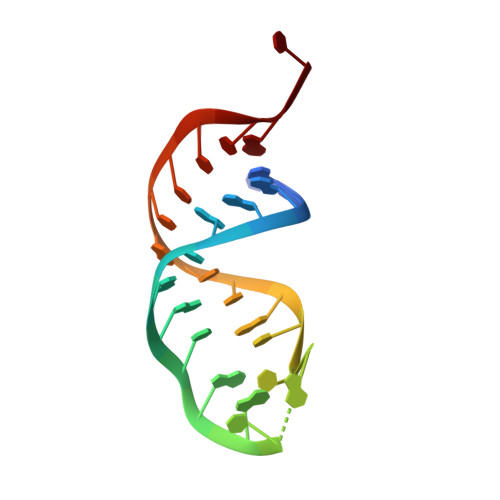
Structural basis for activity switching in polymerases determining the fate of let-7 pre-miRNAs.
Yi, G., Ye, M., Carrique, L., El-Sagheer, A., Brown, T., Norbury, C.J., Zhang, P., Gilbert, R.J.C.(2024) Nat Struct Mol Biol 31: 1426-1438
- PubMed: 39054354 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-024-01357-9
- Primary Citation Related Structures:
8OEF, 8OPP, 8OPS, 8OPT, 8OST - PubMed Abstract:
Tumor-suppressor let-7 pre-microRNAs (miRNAs) are regulated by terminal uridylyltransferases TUT7 and TUT4 that either promote let-7 maturation by adding a single uridine nucleotide to the pre-miRNA 3' end or mark them for degradation by the addition of multiple uridines. Oligo-uridylation is increased in cells by enhanced TUT7/4 expression and especially by the RNA-binding pluripotency factor LIN28A. Using cryogenic electron microscopy, we captured high-resolution structures of active forms of TUT7 alone, of TUT7 plus pre-miRNA and of both TUT7 and TUT4 bound with pre-miRNA and LIN28A. Our structures reveal that pre-miRNAs engage the enzymes in fundamentally different ways depending on the presence of LIN28A, which clamps them onto the TUTs to enable processive 3' oligo-uridylation. This study reveals the molecular basis for mono- versus oligo-uridylation by TUT7/4, as determined by the presence of LIN28A, and thus their mechanism of action in the regulation of cell fate and in cancer.
- Division of Structural Biology, Centre for Human Genetics, Nuffield Department of Medicine, University of Oxford, Oxford, UK.
Organizational Affiliation: