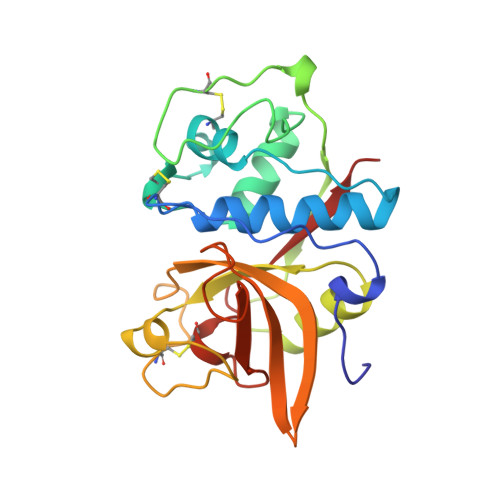
Crystal structure of porcine cathepsin H determined at 2.1 A resolution: location of the mini-chain C-terminal carboxyl group defines cathepsin H aminopeptidase function.
Guncar, G., Podobnik, M., Pungercar, J., Strukelj, B., Turk, V., Turk, D.(1998) Structure 6: 51-61
- PubMed: 9493267 Search on PubMed
- DOI: https://doi.org/10.1016/s0969-2126(98)00007-0
- Primary Citation Related Structures:
8PCH - PubMed Abstract:
Cathepsin H is a lysosomal cysteine protease, involved in intracellular protein degradation. It is the only known mono-aminopeptidase in the papain-like family and is reported to be involved in tumor metastasis. The cathepsin H structure was determined in order to investigate the structural basis for its aminopeptidase activity and thus to provide the basis for structure-based design of synthetic inhibitors. The crystal structure of native porcine cathepsin H was determined at 2.1 A resolution. The structure has the typical papain-family fold. The so-called mini-chain, the octapeptide EPQNCSAT, is attached via a disulfide bond to the body of the enzyme and bound in a narrowed active-site cleft, in the substrate-binding direction. The mini-chain fills the region that in related enzymes comprises the non-primed substrate-binding sites from S2 backwards. The crystal structure of cathepsin H reveals that the mini-chain has a definitive role in substrate recognition and that carbohydrate residues attached to the body of the enzyme are involved in positioning the mini-chain in the active-site cleft. Modeling of a substrate into the active-site cleft suggests that the negatively charged carboxyl group of the C terminus of the mini-chain acts as an anchor for the positively charged N-terminal amino group of a substrate. The observed displacements of the residues within the active-site cleft from their equivalent positions in the papain-like endopeptidases suggest that they form the structural basis for the positioning of both the mini-chain and the substrate, resulting in exopeptidase activity.
- Department of Biochemistry and Molecular Biology, Jozef Stefan Institute, Ljubljana, Slovenia. gregor.guncar@ijs.si
Organizational Affiliation: