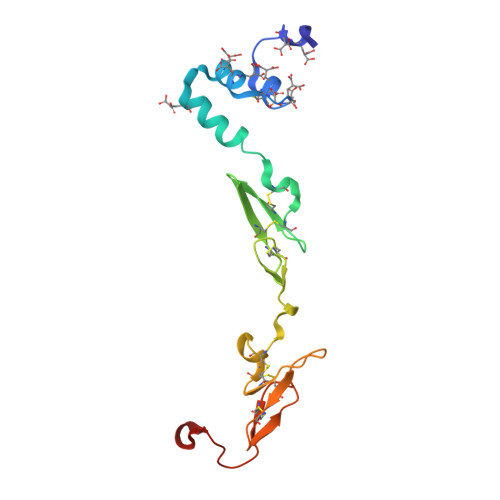
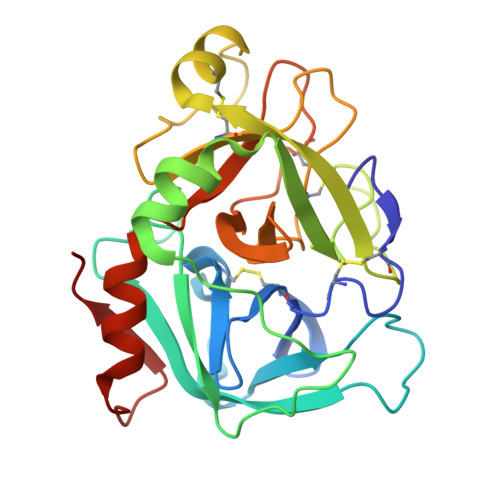
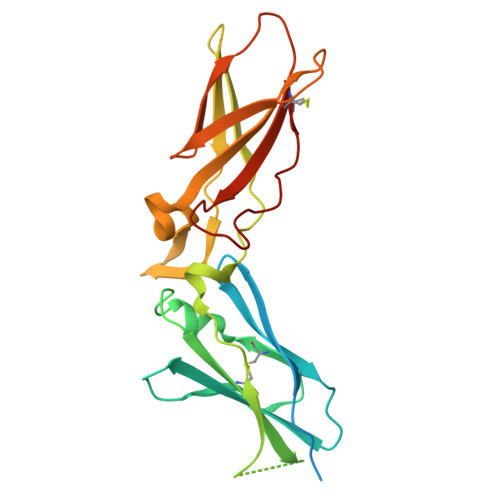
Crystallography and molecular simulations capture S1 pocket collapse in allosteric regulation of factor VIIa and other serine proteases.
Tesmer, L., Hans Matter, H., Klingler, O., Schudok, M., Hessler, G., Mehdipour, A.R., Guessregen, S., Hummer, G., Schreuder, H.A.To be published.