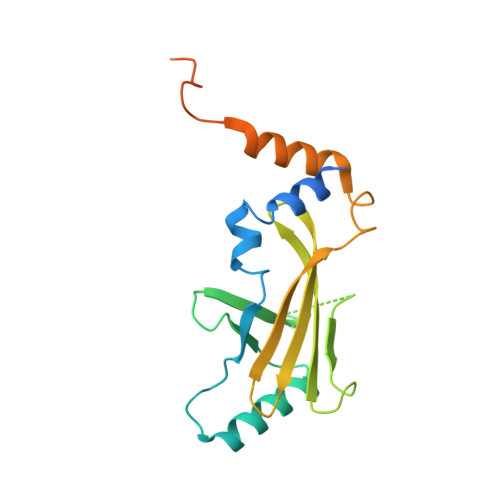
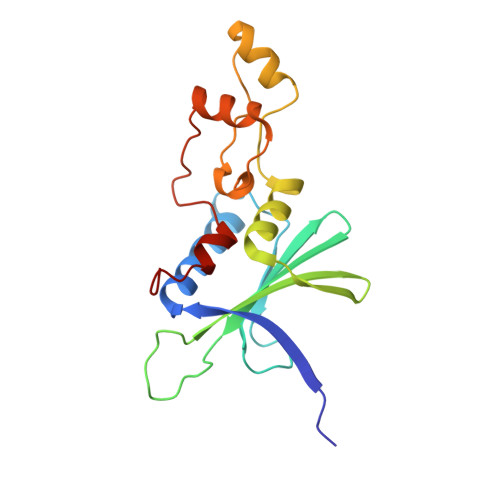
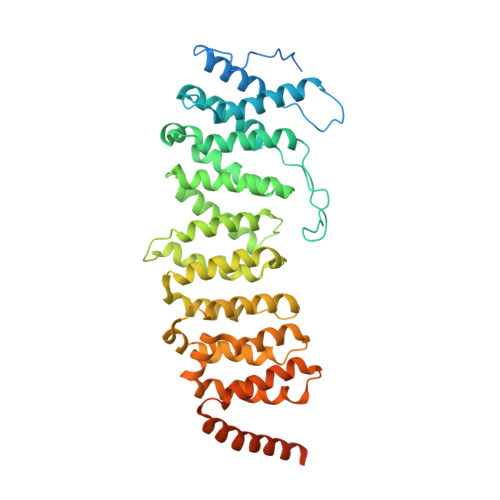
Structural insights into PPP2R5A degradation by HIV-1 Vif.
Hu, Y., Delviks-Frankenberry, K.A., Wu, C., Arizaga, F., Pathak, V.K., Xiong, Y.(2024) Nat Struct Mol Biol 31: 1492-1501
- PubMed: 38789685 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41594-024-01314-6
- Primary Citation Related Structures:
8SZK - PubMed Abstract:
HIV-1 Vif recruits host cullin-RING-E3 ubiquitin ligase and CBFβ to degrade the cellular APOBEC3 antiviral proteins through diverse interactions. Recent evidence has shown that Vif also degrades the regulatory subunits PPP2R5(A-E) of cellular protein phosphatase 2A to induce G2/M cell cycle arrest. As PPP2R5 proteins bear no functional or structural resemblance to A3s, it is unclear how Vif can recognize different sets of proteins. Here we report the cryogenic-electron microscopy structure of PPP2R5A in complex with HIV-1 Vif-CBFβ-elongin B-elongin C at 3.58 Å resolution. The structure shows PPP2R5A binds across the Vif molecule, with biochemical and cellular studies confirming a distinct Vif-PPP2R5A interface that partially overlaps with those for A3s. Vif also blocks a canonical PPP2R5A substrate-binding site, indicating that it suppresses the phosphatase activities through both degradation-dependent and degradation-independent mechanisms. Our work identifies critical Vif motifs regulating the recognition of diverse A3 and PPP2R5A substrates, whereby disruption of these host-virus protein interactions could serve as potential targets for HIV-1 therapeutics.
- Department of Molecular Biophysics and Biochemistry, Yale University, New Haven, CT, USA.
Organizational Affiliation: