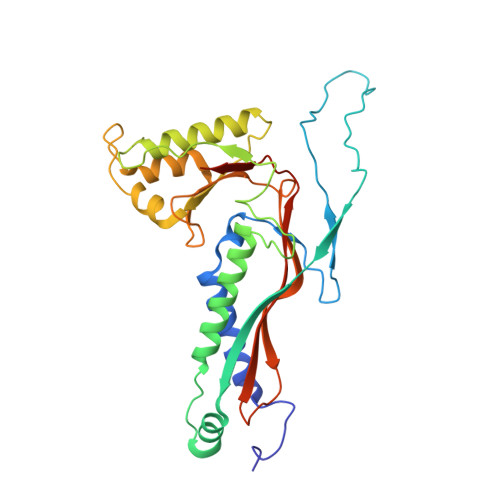
Structure and heterogeneity of a highly cargo-loaded encapsulin shell.
Kwon, S., Andreas, M.P., Giessen, T.W.(2023) J Struct Biol 215: 108022-108022
- PubMed: 37657675
- DOI: https://doi.org/10.1016/j.jsb.2023.108022
- Primary Citation of Related Structures:
8TK7 - PubMed Abstract:
Encapsulins are self-assembling protein nanocompartments able to selectively encapsulate dedicated cargo enzymes. Encapsulins are widespread across bacterial and archaeal phyla and are involved in oxidative stress resistance, iron storage, and sulfur metabolism. Encapsulin shells exhibit icosahedral geometry and consist of 60, 180, or 240 identical protein subunits. Cargo encapsulation is mediated by the specific interaction of targeting peptides or domains, found in all cargo proteins, with the interior surface of the encapsulin shell during shell self-assembly. Here, we report the 2.53 Å cryo-EM structure of a heterologously produced and highly cargo-loaded T3 encapsulin shell from Myxococcus xanthus and explore the systems' structural heterogeneity. We find that exceedingly high cargo loading results in the formation of substantial amounts of distorted and aberrant shells, likely caused by a combination of unfavorable steric clashes of cargo proteins and shell conformational changes. Based on our cryo-EM structure, we determine and analyze the targeting peptide-shell binding mode. We find that both ionic and hydrophobic interactions mediate targeting peptide binding. Our results will guide future attempts at rationally engineering encapsulins for biomedical and biotechnological applications.
- Department of Chemical Engineering, University of Michigan, Ann Arbor, MI 48109, USA.
Organizational Affiliation: