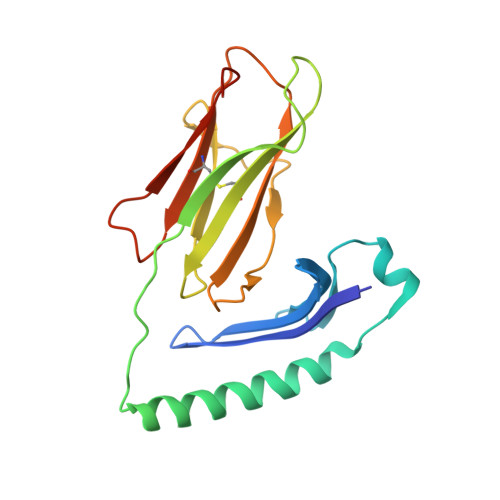
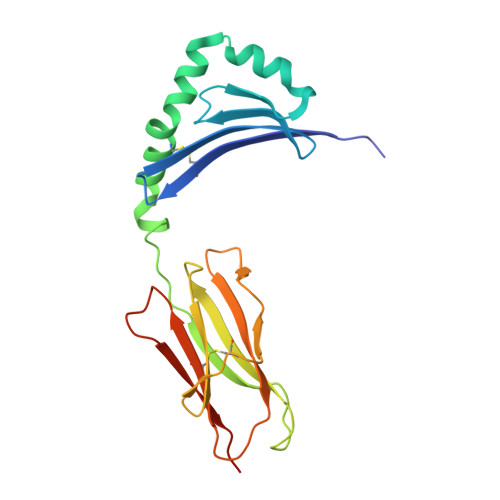
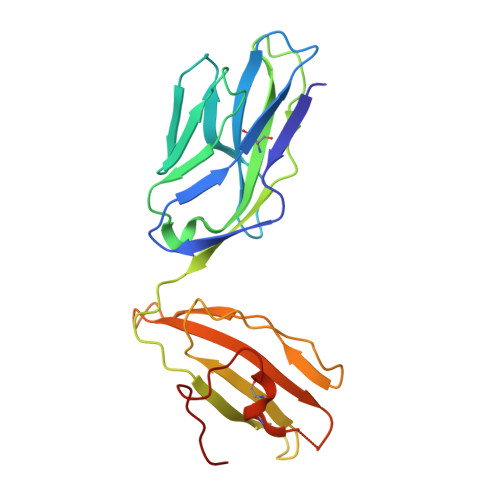
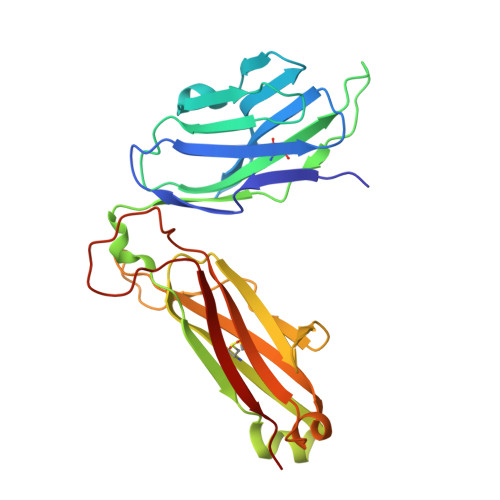
The molecular basis underlying T cell specificity towards citrullinated epitopes presented by HLA-DR4.
Loh, T.J., Lim, J.J., Jones, C.M., Dao, H.T., Tran, M.T., Baker, D.G., La Gruta, N.L., Reid, H.H., Rossjohn, J.(2024) Nat Commun 15: 6201-6201
- PubMed: 39043656 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-024-50511-w
- Primary Citation Related Structures:
8TRL, 8TRQ, 8TRR - PubMed Abstract:
CD4 + T cells recognising citrullinated self-epitopes presented by HLA-DRB1 bearing the shared susceptibility epitope (SE) are implicated in rheumatoid arthritis (RA). However, the underlying T cell receptor (TCR) determinants of epitope specificity towards distinct citrullinated peptide antigens, including vimentin-64cit 59-71 and α-enolase-15cit 10-22 remain unclear. Using HLA-DR4-tetramers, we examine the T cell repertoire in HLA-DR4 transgenic mice and observe biased TRAV6 TCR gene usage across these two citrullinated epitopes which matches with TCR bias previously observed towards the fibrinogen β-74cit 69-81 epitope. Moreover, shared TRAV26-1 gene usage is evident in four α-enolase-15cit 10-22 reactive T cells in three human samples. Crystal structures of mouse TRAV6 + and human TRAV26-1 + TCR-HLA-DR4 complexes presenting vimentin-64cit 59-71 and α-enolase-15cit 10-22 , respectively, show three-way interactions between the TCR, SE, citrulline, and the basis for the biased selection of TRAV genes. Position 2 of the citrullinated epitope is a key determinant underpinning TCR specificity. Accordingly, we provide a molecular basis of TCR specificity towards citrullinated epitopes.
- Infection and Immunity Program and Department of Biochemistry and Molecular Biology, Biomedicine Discovery Institute, Monash University, Clayton, Australia.
Organizational Affiliation: