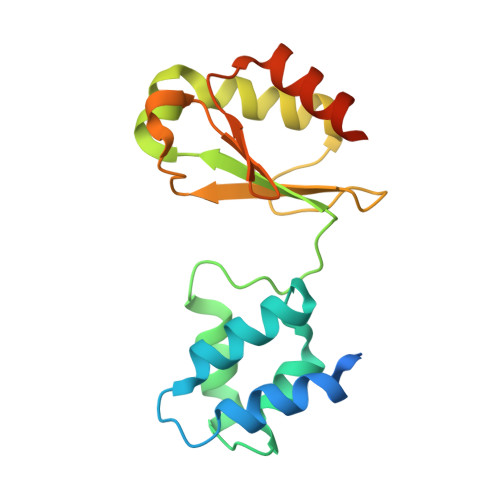
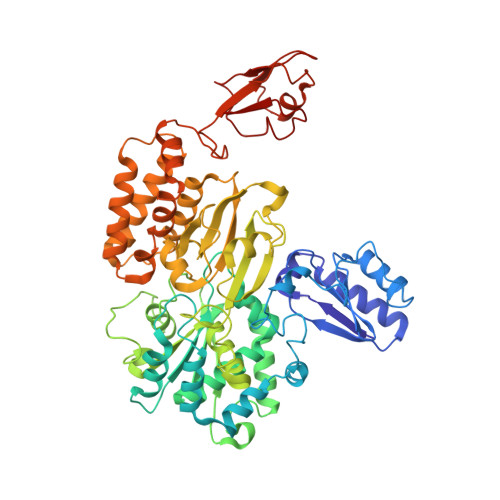
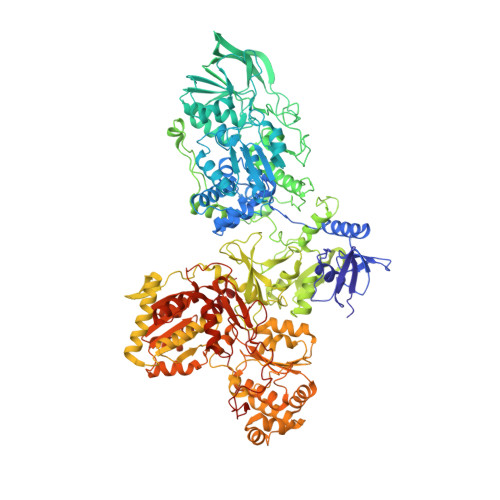
Molecular architecture and electron transfer pathway of the Stn family transhydrogenase.
Kumar, A., Kremp, F., Roth, J., Freibert, S.A., Muller, V., Schuller, J.M.(2023) Nat Commun 14: 5484-5484
- PubMed: 37673911 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-023-41212-x
- Primary Citation Related Structures:
8OH5, 8OH9 - PubMed Abstract:
The challenge of endergonic reduction of NADP + using NADH is overcome by ferredoxin-dependent transhydrogenases that employ electron bifurcation for electron carrier adjustments in the ancient Wood-Ljungdahl pathway. Recently, an electron-bifurcating transhydrogenase with subunit compositions distinct from the well-characterized Nfn-type transhydrogenase was described: the Stn complex. Here, we present the single-particle cryo-EM structure of the Stn family transhydrogenase from the acetogenic bacterium Sporomusa ovata and functionally dissect its electron transfer pathway. Stn forms a tetramer consisting of functional heterotrimeric StnABC complexes. Our findings demonstrate that the StnAB subunits assume the structural and functional role of a bifurcating module, homologous to the HydBC core of the electron-bifurcating HydABC complex. Moreover, StnC contains a NuoG-like domain and a GltD-like NADPH binding domain that resembles the NfnB subunit of the NfnAB complex. However, in contrast to NfnB, StnC lost the ability to bifurcate electrons. Structural comparison allows us to describe how the same fold on one hand evolved bifurcation activity on its own while on the other hand combined with an associated bifurcating module, exemplifying modular evolution in anaerobic metabolism to produce activities critical for survival at the thermodynamic limit of life.
- SYNMIKRO Research Center and Department of Chemistry, Philipps-University of Marburg, Marburg, Germany.
Organizational Affiliation: