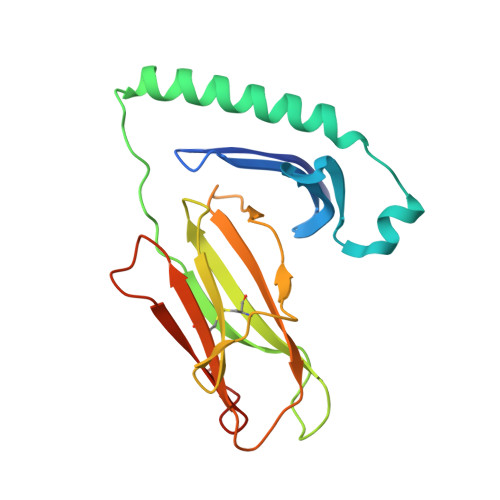
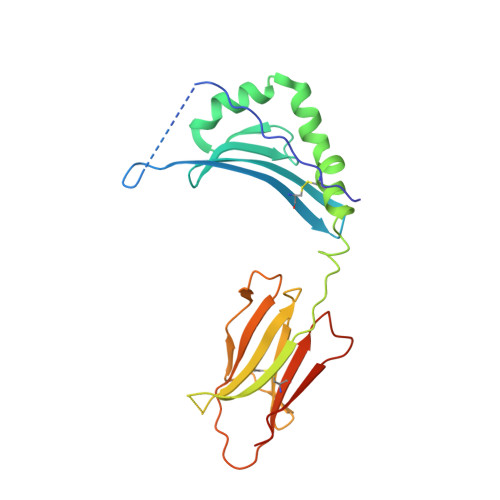
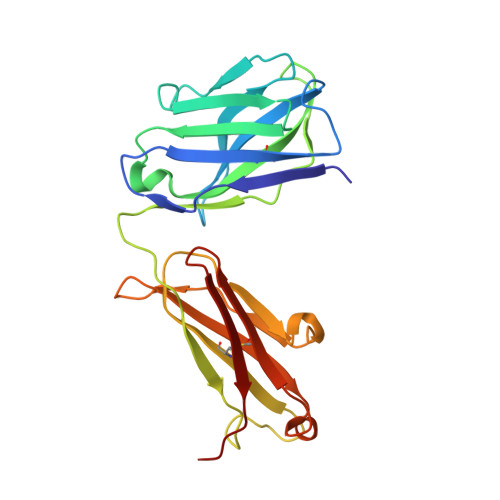
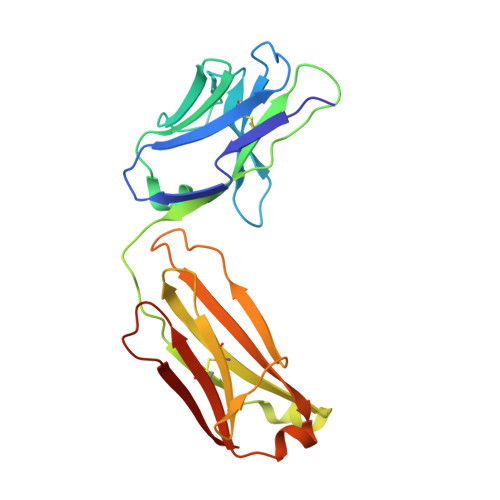
Humanization of Pan-HLA-DR mAb 44H10 Hinges on Critical Residues in the Antibody Framework.
Kassardjian, A., Ivanochko, D., Barber, B., Jetha, A., Julien, J.P.(2024) Antibodies (Basel) 13
- PubMed: 39051333 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.3390/antib13030057
- Primary Citation Related Structures:
9B74, 9B75, 9B76, 9B7B - PubMed Abstract:
Reducing the immunogenicity of animal-derived monoclonal antibodies (mAbs) for use in humans is critical to maximize therapeutic effectiveness and preclude potential adverse events. While traditional humanization methods have primarily focused on grafting antibody Complementarity-Determining Regions (CDRs) on homologous human antibody scaffolds, framework regions can also play essential roles in antigen binding. Here, we describe the humanization of the pan-HLA-DR mAb 44H10, a murine antibody displaying significant involvement of the framework region in antigen binding. Using a structure-guided approach, we identify and restore framework residues that directly interact with the antigen or indirectly modulate antigen binding by shaping the antibody paratope and engineer a humanized antibody with affinity, biophysical profile, and molecular binding basis comparable to that of the parental 44H10 mAb. As a humanized molecule, this antibody holds promise as a scaffold for the development of MHC class II-targeting therapeutics and vaccines.
- Program in Molecular Medicine, The Hospital for Sick Children Research Institute, 686 Bay Street, Toronto, ON M5G 0A4, Canada.
Organizational Affiliation: