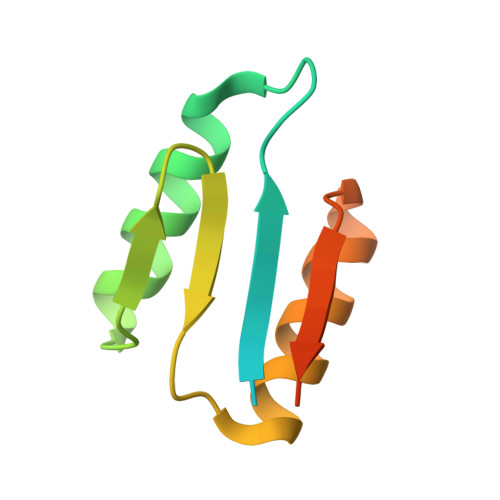
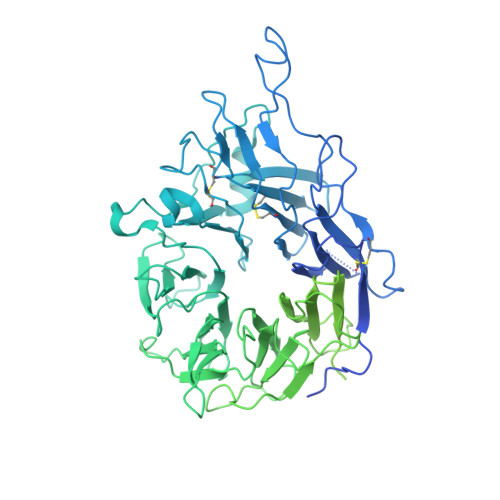
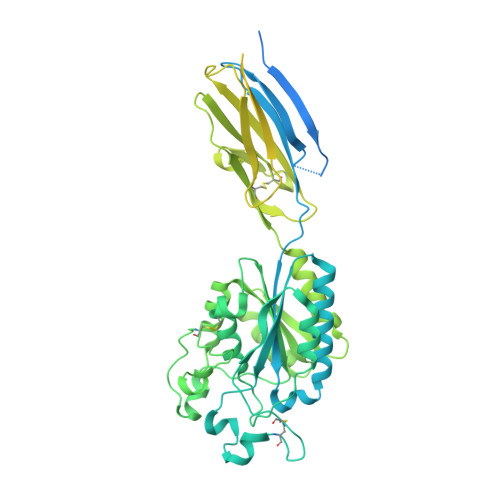
De Novo Design of Integrin alpha 5 beta 1 Modulating Proteins to Enhance Biomaterial Properties.
Wang, X., Guillem-Marti, J., Kumar, S., Lee, D.S., Cabrerizo-Aguado, D., Werther, R., Alamo, K.A.E., Zhao, Y.T., Nguyen, A., Kopyeva, I., Huang, B., Li, J., Hao, Y., Li, X., Brizuela-Velasco, A., Murray, A., Gerben, S., Roy, A., DeForest, C.A., Springer, T., Ruohola-Baker, H., Cooper, J.A., Campbell, M.G., Manero, J.M., Ginebra, M.P., Baker, D.(2025) Adv Mater 37: e2500872-e2500872
- PubMed: 40489013 Search on PubMed
- DOI: https://doi.org/10.1002/adma.202500872
- Primary Citation Related Structures:
9CKV, 9DIA, 9EF2 - PubMed Abstract:
Integrin α5β1 is crucial for cell attachment and migration in development and tissue regeneration, and α5β1 binding proteins can have considerable utility in regenerative medicine and next-generation therapeutics. We use computational protein design to create de novo α5β1-specific modulating miniprotein binders, called NeoNectins, that bind to and stabilize the open state of α5β1. When immobilized onto titanium surfaces and throughout 3D hydrogels, the NeoNectins outperform native fibronectin (FN) and RGD peptides in enhancing cell attachment and spreading, and NeoNectin-grafted titanium implants outperformed FN- and RGD-grafted implants in animal models in promoting tissue integration and bone growth. NeoNectins should be broadly applicable for tissue engineering and biomedicine.
- Department of Biochemistry, University of Washington, Seattle, WA, USA.
Organizational Affiliation: