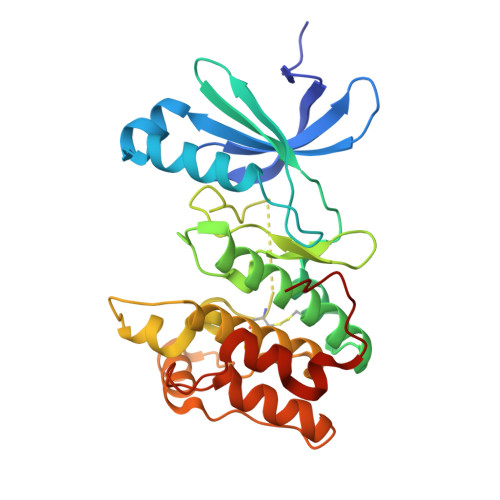
Mechanism and cellular actions of the potent AMPK inhibitor BAY-3827.
Fraguas Bringas, C., Ahangar, M.S., Cuenco, J., Liu, H., Addinsall, A.B., Lindahl, M., Ovens, A.J., Febbraio, M.A., Foretz, M., Goransson, O., Scott, J.W., Zeqiraj, E., Sakamoto, K.(2025) Sci Adv 11: eadx2434-eadx2434
- PubMed: 40845097 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.adx2434
- Primary Citation Related Structures:
9IC2 - PubMed Abstract:
Inhibition of adenosine 5'-monophosphate (AMP)-activated protein kinase (AMPK) is under increasing investigation for its therapeutic potential in many diseases. Existing AMPK inhibitors are however limited, with poor selectivity and substantial off-target effects. Here, we provide mechanistic insights and describe the cellular selectivity of the recently identified AMPK inhibitor BAY-3827. A 2.5-Å cocrystal structure of the AMPK kinase domain with BAY-3827 revealed distinct features including a disulfide bridge between the αD helix Cys 106 and the activation loop residue Cys 174 . This bridge appears to stabilize the activation loop such that Asn 162 repositions the Asp-Phe-Gly (DFG) motif Phe 158 toward the C-terminal lobe, displacing His 137 and disrupting the regulatory spine, promoting an inactive kinase state. In hepatocytes, BAY-3827 blocked AMPK activator (MK-8722)-mediated phosphorylation of ACC1 and corresponding inhibition of lipogenesis. Transcriptome analysis revealed that BAY-3827 down-regulated ~30% of MK-8722-stimulated AMPK-dependent genes. We establish the molecular and cellular basis of BAY-3827's selectivity and utility for delineating AMPK functions while highlighting its limitations.
- Novo Nordisk Foundation Center for Basic Metabolic Research, Faculty of Health and Medical Sciences, University of Copenhagen, Copenhagen 2200, Denmark.
Organizational Affiliation: