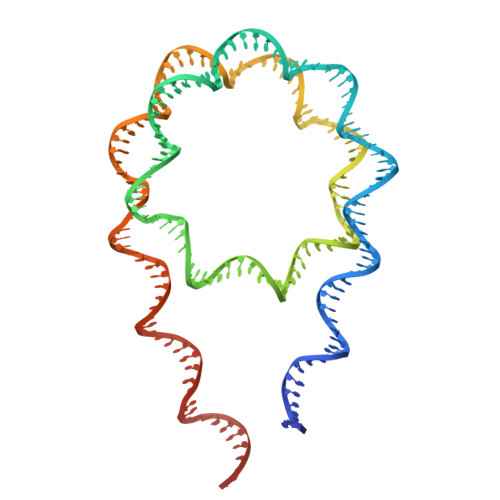
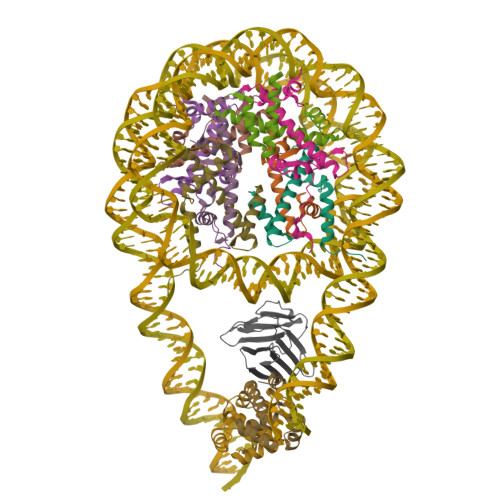
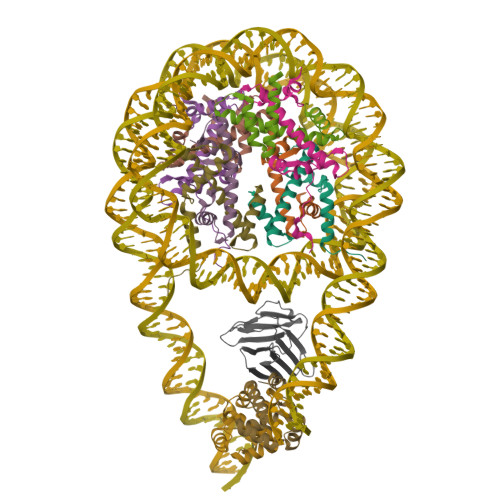
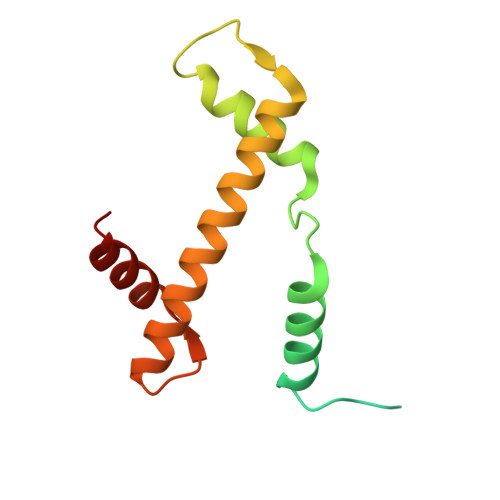
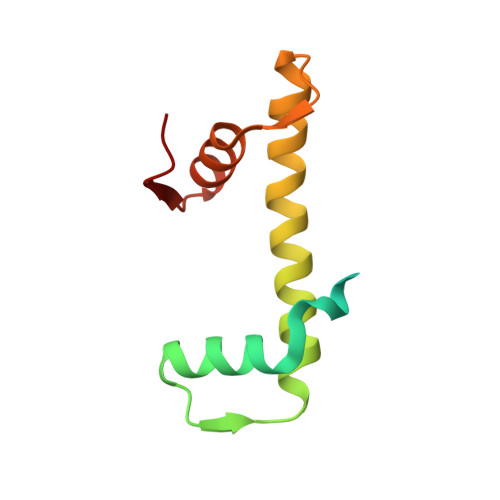
Cryo-EM structures of the BAF-Lamin A/C complex bound to nucleosomes.
Horikoshi, N., Miyake, R., Sogawa-Fujiwara, C., Ogasawara, M., Takizawa, Y., Kurumizaka, H.(2025) Nat Commun 16: 1495-1495
- PubMed: 39929866
- DOI: https://doi.org/10.1038/s41467-025-56823-9
- Primary Citation of Related Structures:
9J8M, 9J8N, 9J8O - PubMed Abstract:
Barrier-to-autointegration factor (BAF) associates with mitotic chromosomes and promotes nuclear envelope assembly by recruiting proteins, such as Lamins, required for the reconstruction of the nuclear envelope and lamina. BAF also mediates chromatin anchoring to the nuclear lamina via Lamin A/C. However, the mechanism by which BAF and Lamin A/C bind chromatin and affect the chromatin organization remains elusive. Here we report the cryo-electron microscopy structures of BAF-Lamin A/C-nucleosome complexes. We find that the BAF dimer complexed with the Lamin A/C IgF domain occupies the nucleosomal dyad position, forming a tripartite nucleosomal DNA binding structure. We also show that the Lamin A/C Lys486 and His506 residues, which are reportedly mutated in lipodystrophy patients, directly contact the DNA at the nucleosomal dyad. Excess BAF-Lamin A/C complexes symmetrically bind other nucleosomal DNA sites and connect two BAF-Lamin A/C-nucleosome complexes. Although the linker histone H1 competes with BAF-Lamin A/C binding at the nucleosomal dyad region, the two BAF-Lamin A/C molecules still bridge two nucleosomes. These findings provide insights into the mechanism by which BAF, Lamin A/C, and/or histone H1 bind nucleosomes and influence chromatin organization within the nucleus.
Organizational Affiliation:
Laboratory of Chromatin Structure and Function, Institute for Quantitative Biosciences, The University of Tokyo, 1-1-1 Yayoi, Bunkyo-ku, Tokyo, Japan.