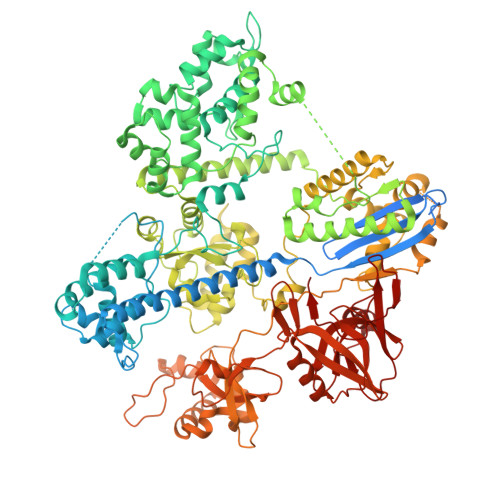
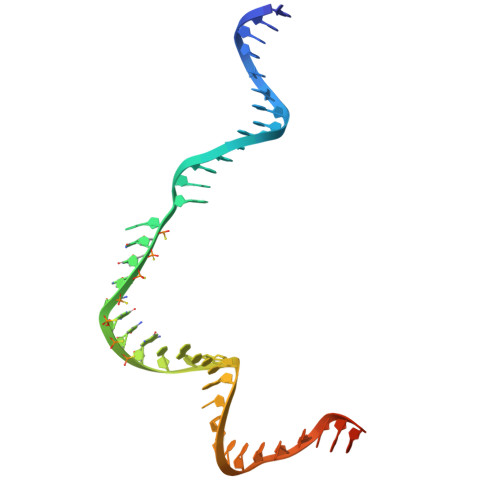
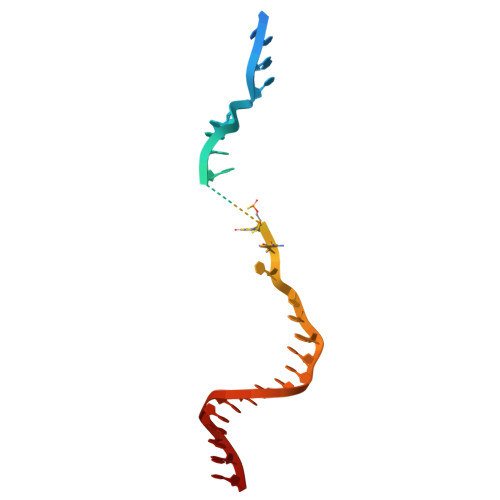
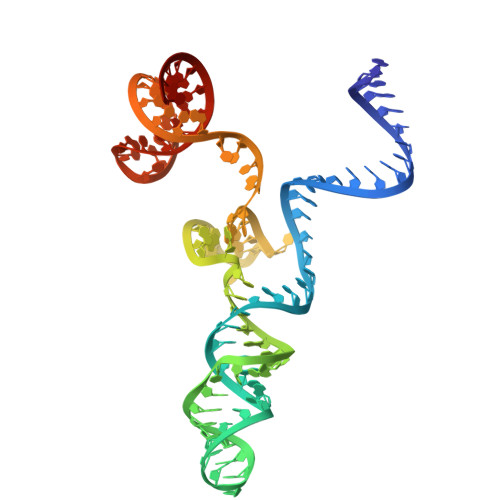
Engineering a compact high-fidelity Staphylococcus aureus Cas9 variant with broader targeting range and mechanistic insights into its activation.
Omura, S.N., Nakagawa, R., Kajimoto, S., Okazaki, S., Ishiguro, S., Mori, H., Onishi, K., Kashiwakura, Y., Hiramoto, T., Horinaka, K., Tanaka, M., Hirano, H., Jividen, K., Yamashita, K., Tsai, S.Q., Yachie, N., Ohmori, T., Nishimasu, H., Nureki, O.(2026) Nat Commun 17
- PubMed: 41991526 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-026-71626-2
- Primary Citation Related Structures:
8ZCY, 8ZCZ, 8ZD0, 8ZDA, 9MB6, 9MB7 - PubMed Abstract:
Staphylococcus aureus Cas9 (SaCas9) is smaller than the widely used Streptococcus pyogenes Cas9 (SpCas9) and has been harnessed for gene therapy using an adeno-associated virus vector. However, SaCas9 requires a longer NNGRRT (where N is any nucleotide and R is A or G) protospacer adjacent motif (PAM) for target DNA recognition, thereby restricting the targeting range. Although PAM-relaxed Cas9 variants have been developed, expanded targeting is often accompanied by compromised target specificity. Here, we report the rational engineering of eSaCas9-NNG, a SaCas9 variant that recognizes relaxed NNG PAMs while maintaining high target fidelity, thereby overcoming a fundamental trade-off in Cas9-based genome editing. eSaCas9-NNG efficiently induces indels and base conversions at endogenous sites bearing NNG PAMs in human cells and mice, with editing efficiencies comparable to those of other PAM-relaxed nucleases, including SpRY, SpG, and iGeoCas9, but with reduced off-target activity. We further determine the cryo-electron microscopy structures of eSaCas9-NNG in five distinct functional states, revealing the structural basis for its relaxed PAM recognition, improved target specificity, and nuclease activation. Overall, our findings demonstrate that eSaCas9-NNG could be used as a versatile genome editing tool for in vivo gene therapy, and improve our mechanistic understanding of the diverse CRISPR-Cas9 nucleases.
- Department of Biological Sciences, Graduate School of Science, The University of Tokyo, Tokyo, Japan.
Organizational Affiliation: