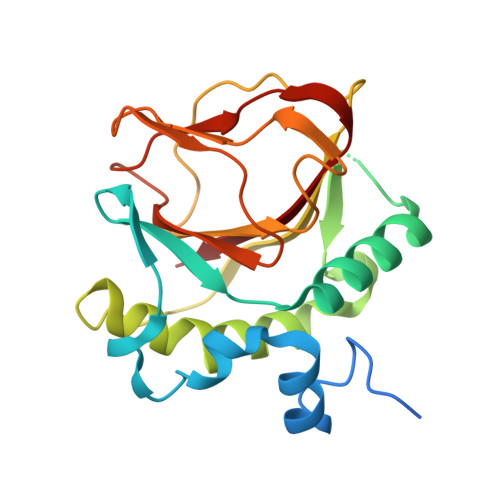
Structure of an Fe 2+ -binding-deficient mimiviral collagen lysyl hydroxylase.
Chen, T., Buhlheller, C., Guo, H.(2025) Acta Crystallogr F Struct Biol Commun 81: 235-240
- PubMed: 40314237 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1107/S2053230X25003735
- Primary Citation Related Structures:
9NEV - PubMed Abstract:
Collagen lysyl hydroxylases catalyze the hydroxylation of collagen lysine residues during collagen synthesis in animals and mimiviruses. Lysyl hydroxylation is crucial for collagen fibrogenesis and function. We previously demonstrated that recombinant mimiviral and human collagen lysyl hydroxylases, isolated from bacterial and mammalian cells, have Fe 2+ in their active sites, suggesting that lysyl hydroxylases have a high affinity for Fe 2+ . We found that Fe 2+ binding stabilizes lysyl hydroxylase dimers, although the underlying mechanism remains unclear. Crystal structure analysis of mimiviral lysyl hydroxylase revealed that Fe 2+ is coordinated by a 2His-1Asp (His825/His877/Asp827) triad, with a nearby highly conserved histidine residue (His869) involved in an alternative 2His-1Asp triad (His869/His877/Asp827). This unique structural architecture suggests that the alternative 2His-1Asp triad may also bind Fe 2+ . To investigate whether the alternative 2His-1Asp triad binds Fe 2+ and how Fe 2+ binding regulates lysyl hydroxylase dimerization, we crystallized the mimiviral lysyl hydroxylase mutant His825Ala, which lacks one 2His-1Asp (His825/His877/Asp827) triad but retains the alternative triad (His869/His877/Asp827). Despite providing Fe 2+ during crystallization, we found no electron density near the alternative 2His-1Asp triad in the His825Ala mutant, indicating that the alternative 2His-1Asp triad does not bind Fe 2+ with high affinity. Although the His825Ala mutant forms a dimer similar to the wild-type enzyme, conformational changes occur in residues near Ala825, including Leu873, which is critical for dimerization. These structural findings provide new insights into the function and regulation of collagen lysyl hydroxylases.
- Department of Molecular and Cellular Biochemistry, Markey Cancer Center, University of Kentucky, 741 South Limestone Avenue, Lexington, KY 40536-0509, USA.
Organizational Affiliation: