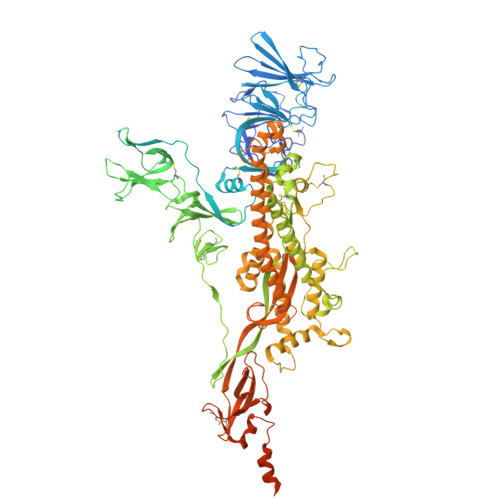
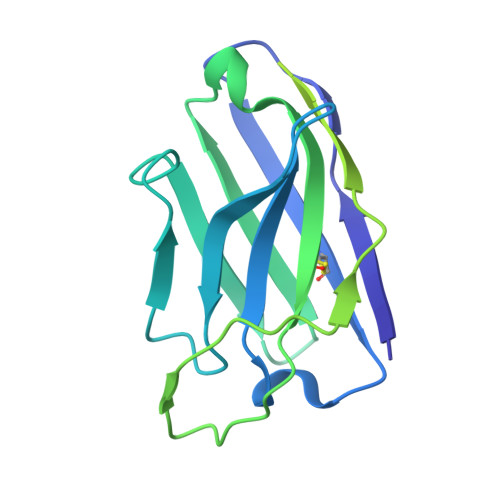
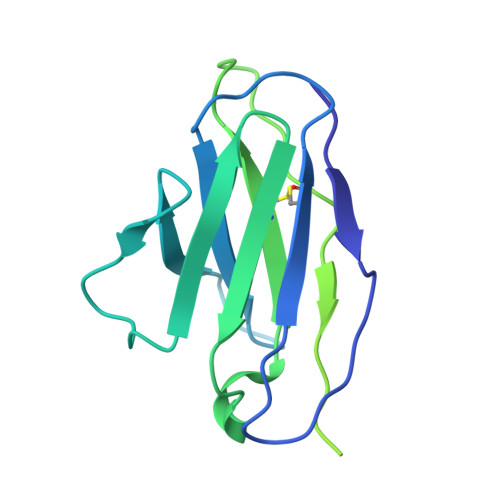
A neutralizing human antibody induces movement of the HCoV-229E receptor binding domain.
Liccione, M.F., Scobey, T., Froggatt, H.M., Pajon, C., Yount, B.L., Yin, Q., Spence, T.N., Walter, E.B., Saunders, K.O., Edwards, R.J., Baric, R.S., Haynes, B.F., Cain, D.W., Sheahan, T.P., Wrapp, D.(2025) Commun Biol 9: 168-168
- PubMed: 41476125
- DOI: https://doi.org/10.1038/s42003-025-09446-1
- Primary Citation of Related Structures:
9OFO, 9OFP, 9OFQ - PubMed Abstract:
HCoV-229E is an endemic Alphacoronavirus that typically causes common cold-like disease in most healthy adults, but can also cause severe respiratory disease in the very young and the elderly. Although the virus was discovered over sixty years ago and undergoes continuous antigenic drift, remarkably little is known about the humoral immune response to HCoV-229E infection. Here we report the isolation of two receptor binding domain-targeting neutralizing human antibodies raised in response to natural HCoV-229E infection. One of these, DH1533, potently neutralizes HCoV-229E, binds to spike with sub-nanomolar affinity and prevents the association between the RBD and the host cell receptor aminopeptidase N. Structural characterization of this antibody bound to HCoV-229E spike delineated a neutralization-sensitive epitope on the RBD and revealed that DH1533 induces conformational flexibility in neighboring RBDs, reminiscent of the "up-and-down" kinetics observed in the related Betacoronavirus spikes. These findings provide insight into the humoral immune response to HCoV-229E infection and will serve as a guide for the design of future therapeutic interventions.
- Department of Epidemiology, University of North Carolina at Chapel Hill, Chapel Hill, NC, USA.
Organizational Affiliation: