A de novo missense variant in MIDEAS results in increased deacetylase activity of the MiDAC HDAC complex causing a neurodevelopmental syndrome.
Fairall, L., Sirvydis, K., Turnbull, R.E., Knottnerus, S.J., Gonchar, O., Muskett, F.W., Jukes-Jones, R., van Brussel, L., van de Geer, E., van Gassen, K., Badenhorst, P., Johnson, D., Terhal, P.A., van Hasselt, P.M., van Jaarsveld, R.H., Schwabe, J.W.(2025) Nat Commun 16: 10472-10472
- PubMed: 41290615
- DOI: https://doi.org/10.1038/s41467-025-65472-x
- Primary Citation of Related Structures:
9R4I - PubMed Abstract:
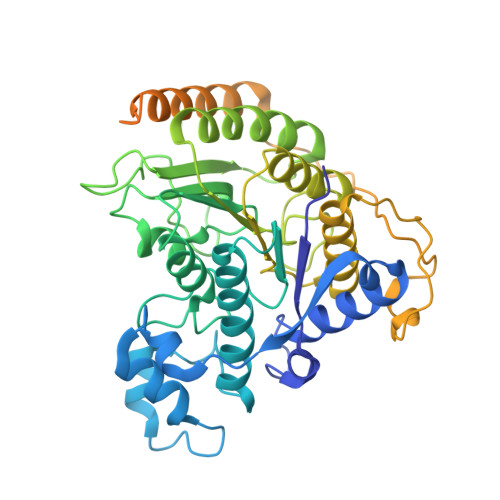
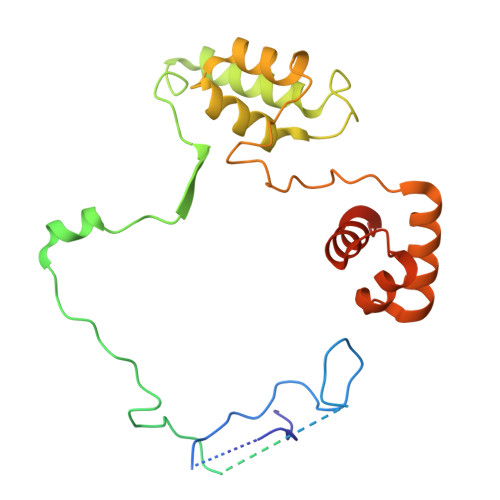
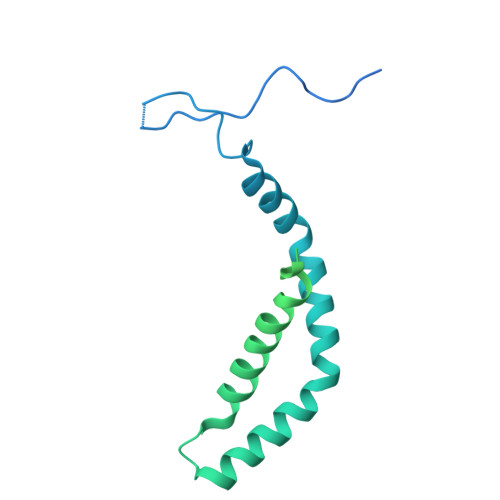
MIDEAS is a scaffold protein that, together with DNTTIP1, mediates assembly of the MiDAC histone deacetylase complex. Mice lacking MiDAC die before birth suggesting a key developmental function. Here, we report two unrelated individuals, with a multisystem disorder characterised by delayed speech development, joint contractures, dysmorphic features and dysmotility of the gut. Both individuals have the same de novo heterozygous missense variant in MIDEAS (p.Tyr654Ser). A cryoEM structure of the MiDAC complex reveals that this amino acid is located in a conserved auto-inhibitory loop that covers the active site of the deacetylase enzyme. We suggest that the variant results in loop displacement leading to elevated deacetylase activity. In support, we observe reciprocal gene expression changes in patient fibroblasts compared with a cell line following rapid MiDAC degradation. Our results establish MIDEAS as a dominant monogenic disease gene and that hyperactivity of the MiDAC complex results in a characteristic multisystem disorder.
- Institute for Structural and Chemical Biology, Department of Molecular and Cell Biology, University of Leicester, Leicester, UK.
Organizational Affiliation: