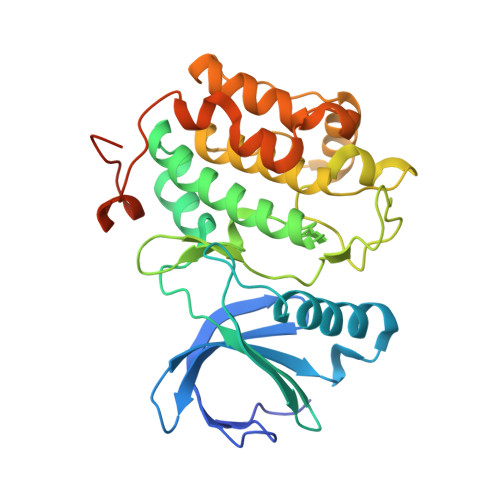
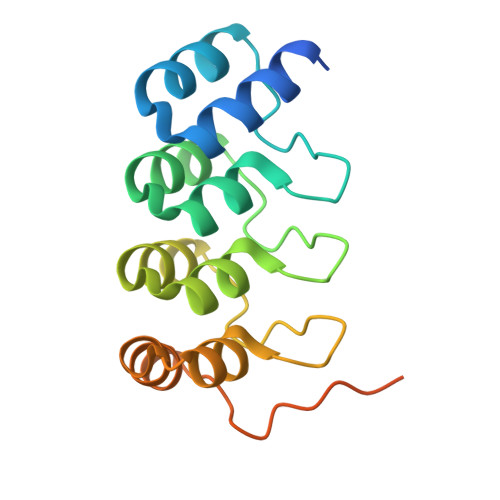
Polo-like kinase 1-inhibitor co-complex structures via the surface-entropy reduction approach and a DARPin-assisted approach.
Eberspaecher, U., Schmitz, A.A., Siemeister, G., Bomer, U., Bandeiras, T.M., Matias, P.M., Schulze, V.K., Hillig, R.C.(2025) Acta Crystallogr D Struct Biol 81: 718-733
- PubMed: 41246821 Search on PubMed
- DOI: https://doi.org/10.1107/S2059798325009325
- Primary Citation Related Structures:
9R1W, 9R1X, 9R1Y, 9R8B, 9R8C - PubMed Abstract:
Polo-like kinase 1 (PLK1) is a major regulator of cell division and has been pursued as a drug target for cancer therapy for a long time. Crystallization of the kinase domain has proven to be exceptionally challenging. Previously, we published a crystallization approach using a PLK1-specific designed ankyrin-repeat protein (DARPin) as a crystallization facilitator. Here, we report an alternative route: crystallization was successful after the introduction of a double mutation which reduced surface entropy and enabled the formation of a new crystal contact. This new PLK1 crystallization system was used to determine the first co-complex crystal structure of the Bayer thiazolidinone lead series, as well as crystal structures with representatives of two competitor inhibitor series. The molecular binding modes of these three inhibitors are analysed and discussed, and the surface-entropy reduction approach is compared with the surface modifications employed by us and others to enable the crystallization of PLK1.
- Research and Development, Pharmaceuticals, Bayer AG, Muellerstrasse 178, 13353 Berlin, Germany.
Organizational Affiliation: