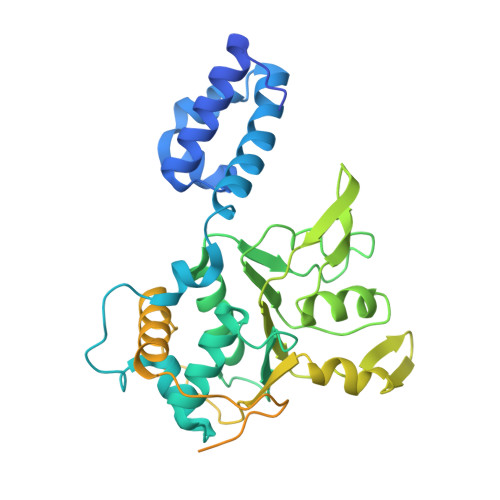
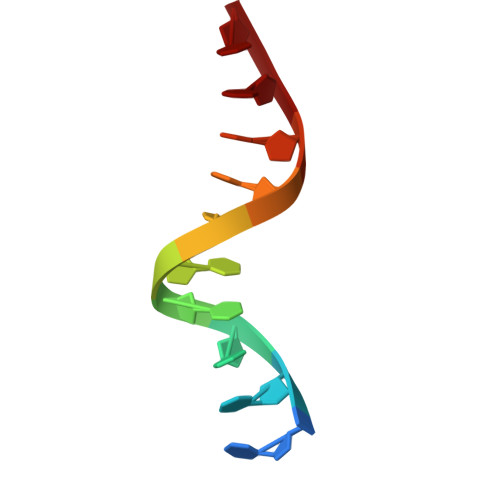
Structural basis for Rep-mediated adeno-associated virus packaging.
Xu, V., McInnes, A., Wake, M., Acebron-Garcia-de-Eulate, M., Barritt, J.D., Bubeck, D., Rouse, S.L.(2026) Cell Rep 45: 117044-117044
- PubMed: 41790558
- DOI: https://doi.org/10.1016/j.celrep.2026.117044
- Primary Citation of Related Structures:
9RM5, 9RQT, 9RRS, 9RWG, 9S0N, 9S10 - PubMed Abstract:
Adeno-associated viruses (AAVs) are parvoviruses utilized as gene therapy vectors. However, the AAV packaging mechanism is unresolved at the molecular level, creating a bottleneck for vector manufacturing, safety, and efficacy. Here, cryo-EM structures of the Rep helicase packaging motor in complex with the packaging marker DNA (ITR) and the Rep-AAV8 capsid complex are presented. Rep-ITR complexes reveal dynamic oligomeric states on the DNA, elucidating the strand separation mechanism coupled to its ATPase cycle. We observe Rep preferentially bound to empty capsids, with a binding interface likely conserved across the virus family. This complex also unveils a cryptic capsid ATP-binding site which, alongside Rep binding, triggers structural rearrangements priming the capsid for packaging. Collectively, these findings advance the understanding of Rep-mediated packaging, with significant implications for parvovirus virology and viral vector design.
- Department of Life Sciences, Imperial College London, London SW7 2AZ, UK; AstraZeneca, Discovery Sciences, BioPharmaceuticals R&D Unit, Cambridge CB2 0AA, UK.
Organizational Affiliation: