Pi-bound ADP-Glucose Pyrophosphorylase
Wu, Y.T., Lin, H.J., Fan, M.R.To be published.
Experimental Data Snapshot
wwPDB Validation 3D Report Full Report
Macromolecule Content
Entity ID: 1 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Glucose-1-phosphate adenylyltransferase small subunit, chloroplastic | 470 | Arabidopsis thaliana | Mutation(s): 0 Gene Names: APS1, ADG1, APS, At5g48300, K23F3.2 EC: 2.7.7.27 | 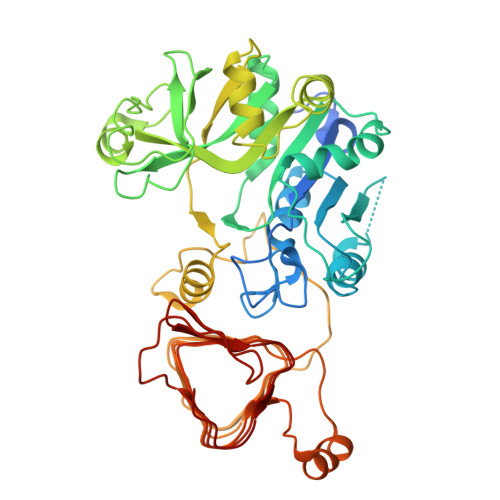 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P55228 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
Entity ID: 2 | |||||
|---|---|---|---|---|---|
| Molecule | Chains | Sequence Length | Organism | Details | Image |
| Glucose-1-phosphate adenylyltransferase large subunit 1, chloroplastic | 477 | Arabidopsis thaliana | Mutation(s): 0 Gene Names: ADG2, APL1, At5g19220, T24G5.120 EC: 2.7.7.27 | 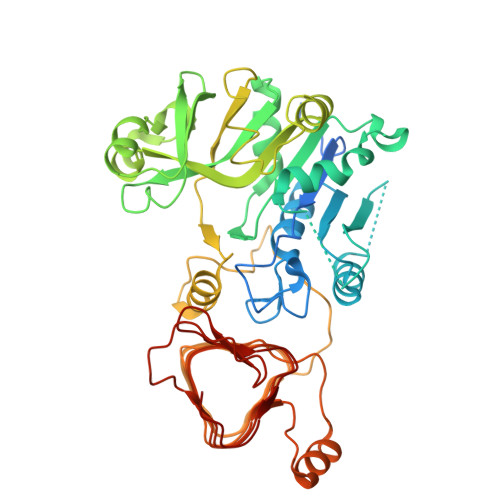 | |
UniProt | |||||
Entity Groups | |||||
| Sequence Clusters | 30% Identity50% Identity70% Identity90% Identity95% Identity100% Identity | ||||
| UniProt Group | P55229 | ||||
Sequence AnnotationsExpand | |||||
Reference Sequence | |||||
| Ligands 1 Unique | |||||
|---|---|---|---|---|---|
| ID | Chains | Name / Formula / InChI Key | 2D Diagram | 3D Interactions | |
| PO4 (Subject of Investigation/LOI) Download:Ideal Coordinates CCD File | E [auth A] F [auth B] G [auth B] H [auth C] I [auth D] | PHOSPHATE ION O4 P NBIIXXVUZAFLBC-UHFFFAOYSA-K |  | ||
| Task | Software Package | Version |
|---|---|---|
| MODEL REFINEMENT | PHENIX | |
| RECONSTRUCTION | cryoSPARC |
| Funding Organization | Location | Grant Number |
|---|---|---|
| Chinese Academy of Sciences | China | XDB0630101 |