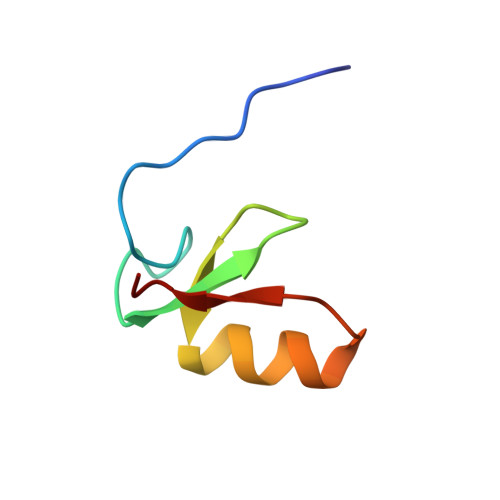
The B-box 1 dimer of human promyelocytic leukemia protein.
Huang, S.Y., Naik, M.T., Chang, C.F., Fang, P.J., Wang, Y.H., Shih, H.M., Huang, T.H.(2014) J Biomol NMR 60: 275-281
- PubMed: 25355412 Search on PubMed
- DOI: https://doi.org/10.1007/s10858-014-9869-4
- Primary Citation Related Structures:
2MVW - Institute of Biomedical Sciences, Academia Sinica, Nankang, Taipei, 11529, Taiwan, ROC.
Organizational Affiliation: