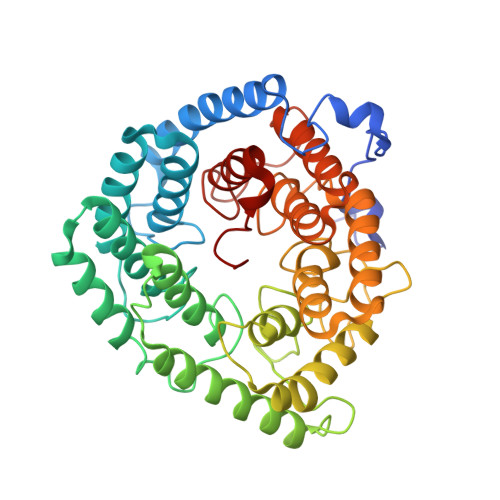
Structure of human lanthionine synthetase C-like protein 1 and its interaction with Eps8 and glutathione
Zhang, W., Wang, L., Liu, Y., Xu, J., Zhu, G., Cang, H., Li, X., Bartlam, M., Hensley, K., Li, G., Rao, Z., Zhang, X.C.(2009) Genes Dev 23: 1387-1392
- PubMed: 19528316 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1101/gad.1789209
- Primary Citation Related Structures:
3E6U, 3E73 - PubMed Abstract:
Eukaryotic lanthionine synthetase C-like protein 1 (LanCL1) is homologous to prokaryotic lanthionine cyclases, yet its biochemical functions remain elusive. We report the crystal structures of human LanCL1, both free of and complexed with glutathione, revealing glutathione binding to a zinc ion at the putative active site formed by conserved GxxG motifs. We also demonstrate by in vitro affinity analysis that LanCL1 binds specifically to the SH3 domain of a signaling protein, Eps8. Importantly, expression of LanCL1 mutants defective in Eps8 interaction inhibits nerve growth factor (NGF)-induced neurite outgrowth, providing evidence for the biological significance of this novel interaction in cellular signaling and differentiation.
- National Laboratory of Biomacromolecules, Institute of Biophysics, Chinese Academy of Sciences, Beijing 100101, China.
Organizational Affiliation: