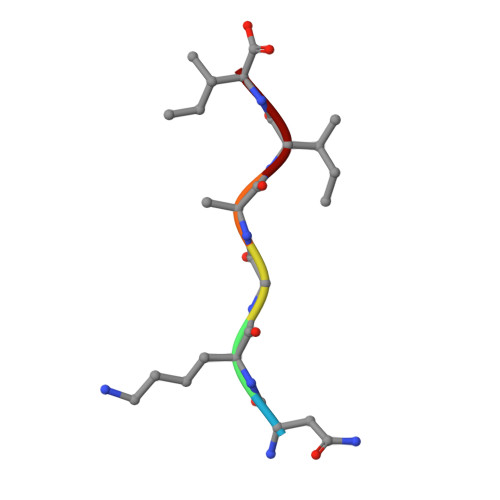
Molecular basis for amyloid-beta polymorphism.
Colletier, J.P., Laganowsky, A., Landau, M., Zhao, M., Soriaga, A.B., Goldschmidt, L., Flot, D., Cascio, D., Sawaya, M.R., Eisenberg, D.(2011) Proc Natl Acad Sci U S A 108: 16938-16943
- PubMed: 21949245 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1112600108
- Primary Citation Related Structures:
2Y29, 2Y2A, 2Y3J, 2Y3K, 2Y3L, 3OW9, 3PZZ, 3Q2X - PubMed Abstract:
Amyloid-beta (Aβ) aggregates are the main constituent of senile plaques, the histological hallmark of Alzheimer's disease. Aβ molecules form β-sheet containing structures that assemble into a variety of polymorphic oligomers, protofibers, and fibers that exhibit a range of lifetimes and cellular toxicities. This polymorphic nature of Aβ has frustrated its biophysical characterization, its structural determination, and our understanding of its pathological mechanism. To elucidate Aβ polymorphism in atomic detail, we determined eight new microcrystal structures of fiber-forming segments of Aβ. These structures, all of short, self-complementing pairs of β-sheets termed steric zippers, reveal a variety of modes of self-association of Aβ. Combining these atomic structures with previous NMR studies allows us to propose several fiber models, offering molecular models for some of the repertoire of polydisperse structures accessible to Aβ. These structures and molecular models contribute fundamental information for understanding Aβ polymorphic nature and pathogenesis.
- Howard Hughes Medical Institute, Department of Biological Chemistry and Chemistry and Biochemistry, UCLA-DOE Institute for Genomics and Proteomics, University of California, Los Angeles, CA 90095, USA.
Organizational Affiliation: