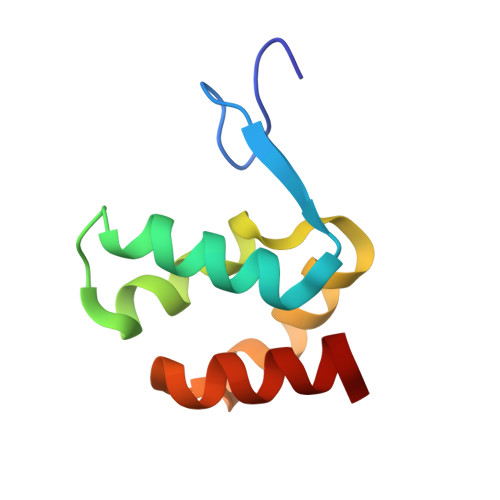
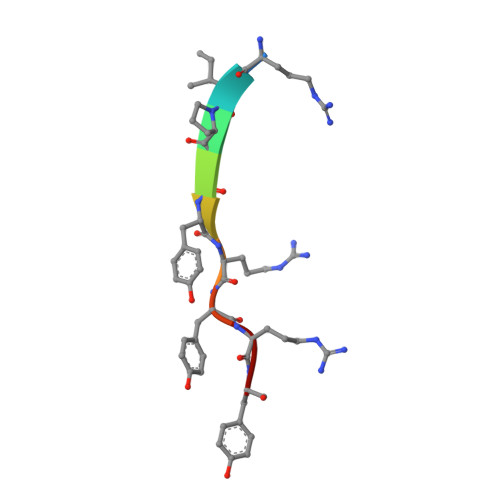
Structural basis of arc binding to synaptic proteins: implications for cognitive disease.
Zhang, W., Wu, J., Ward, M.D., Yang, S., Chuang, Y.A., Xiao, M., Li, R., Leahy, D.J., Worley, P.F.(2015) Neuron 86: 490-500
- PubMed: 25864631 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1016/j.neuron.2015.03.030
- Primary Citation Related Structures:
4X3H, 4X3I, 4X3X - PubMed Abstract:
Arc is a cellular immediate-early gene (IEG) that functions at excitatory synapses and is required for learning and memory. We report crystal structures of Arc subdomains that form a bi-lobar architecture remarkably similar to the capsid domain of human immunodeficiency virus (HIV) gag protein. Analysis indicates Arc originated from the Ty3/Gypsy retrotransposon family and was "domesticated" in higher vertebrates for synaptic functions. The Arc N-terminal lobe evolved a unique hydrophobic pocket that mediates intermolecular binding with synaptic proteins as resolved in complexes with TARPγ2 (Stargazin) and CaMKII peptides and is essential for Arc's synaptic function. A consensus sequence for Arc binding identifies several additional partners that include genes implicated in schizophrenia. Arc N-lobe binding is inhibited by small chemicals suggesting Arc's synaptic action may be druggable. These studies reveal the remarkable evolutionary origin of Arc and provide a structural basis for understanding Arc's contribution to neural plasticity and disease.
- Solomon H. Snyder Department of Neuroscience, Johns Hopkins University School of Medicine, Baltimore, MD 21205, USA.
Organizational Affiliation: