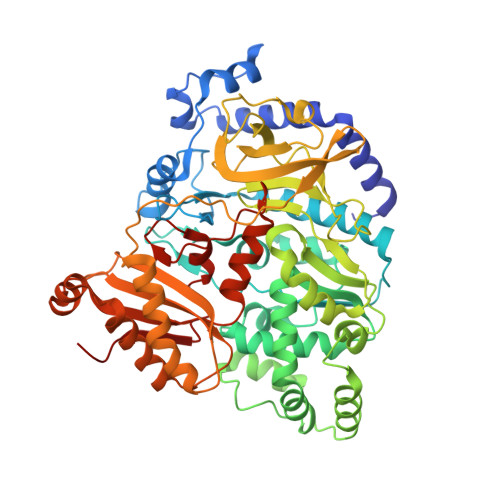
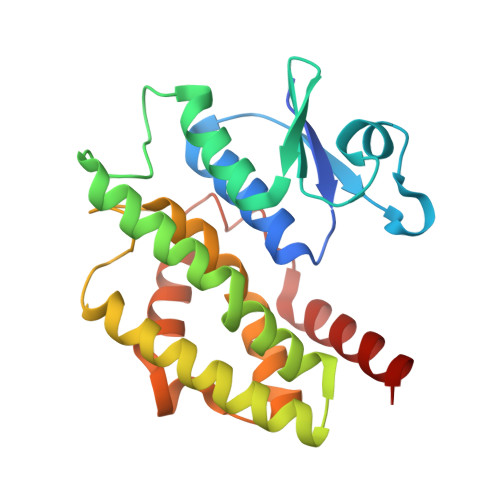
Structural basis of jasmonate-amido synthetase FIN219 in complex with glutathione S-transferase FIP1 during the JA signal regulation
Chen, C.Y., Ho, S.S., Kuo, T.Y., Hsieh, H.L., Cheng, Y.S.(2017) Proc Natl Acad Sci U S A 114: E1815-E1824
- PubMed: 28223489 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1073/pnas.1609980114
- Primary Citation Related Structures:
5ECH, 5ECI, 5ECK, 5ECL, 5ECM, 5ECN, 5ECO, 5ECP, 5ECQ, 5ECR, 5ECS, 5GZZ - PubMed Abstract:
Far-red (FR) light-coupled jasmonate (JA) signaling is necessary for plant defense and development. FR insensitive 219 (FIN219) is a member of the Gretchen Hagen 3 (GH3) family of proteins in Arabidopsis and belongs to the adenylate-forming family of enzymes. It directly controls biosynthesis of jasmonoyl-isoleucine in JA-mediated defense responses and interacts with FIN219-interacting protein 1 (FIP1) under FR light conditions. FIN219 and FIP1 are involved in FR light signaling and are regulators of the interplay between light and JA signaling. However, how their interactions affect plant physiological functions remains unclear. Here, we demonstrate the crystal structures of FIN219-FIP1 while binding with substrates at atomic resolution. Our results show an unexpected FIN219 conformation and demonstrate various differences between this protein and other members of the GH3 family. We show that the rotated C-terminal domain of FIN219 alters ATP binding and the core structure of the active site. We further demonstrate that this unique FIN219-FIP1 structure is crucial for increasing FIN219 activity and determines the priority of substrate binding. We suggest that the increased FIN219 activity resulting from the complex form, a conformation for domain switching, allows FIN219 to switch to its high-affinity mode and thereby enhances JA signaling under continuous FR light conditions.
- Institute of Plant Biology, National Taiwan University, Taipei 10617, Taiwan.
Organizational Affiliation: