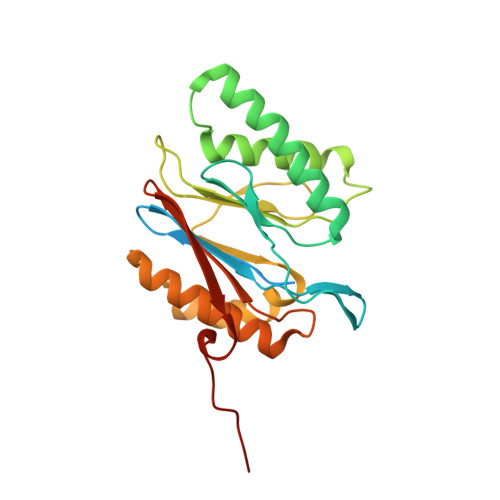
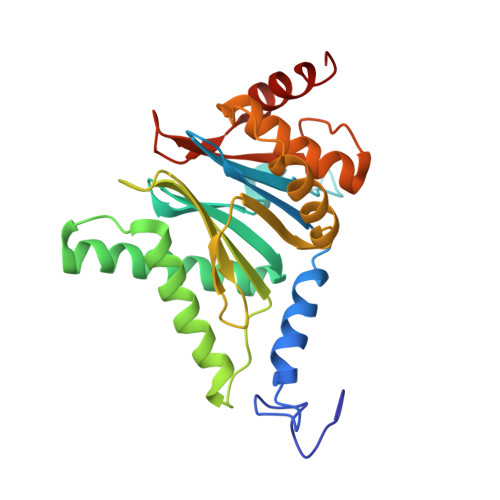
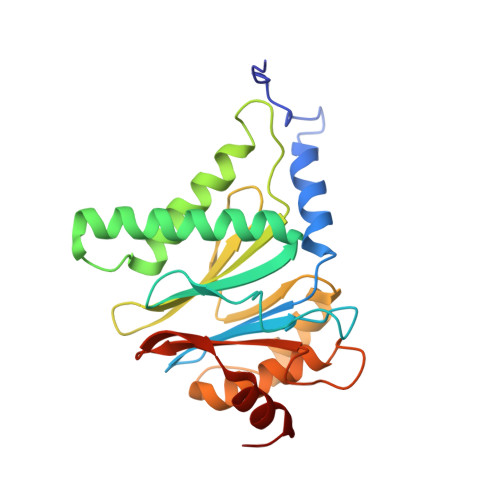
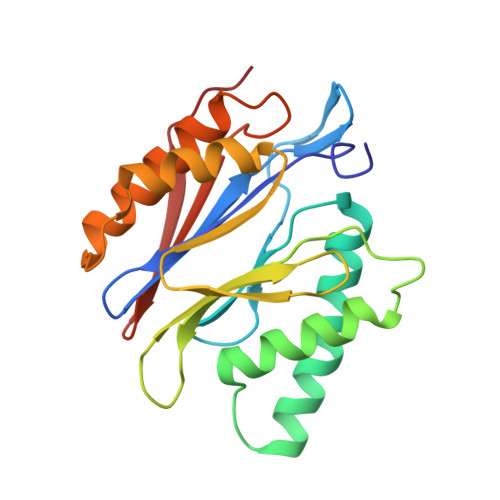
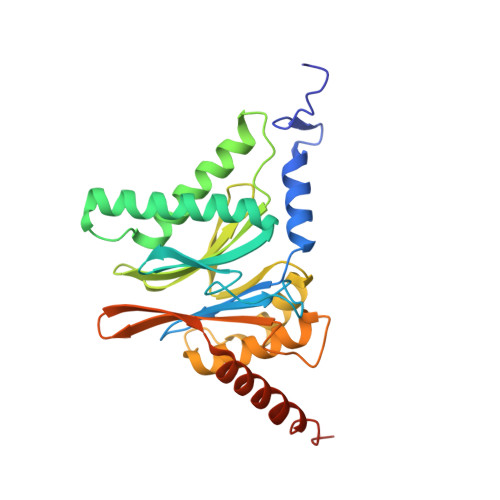
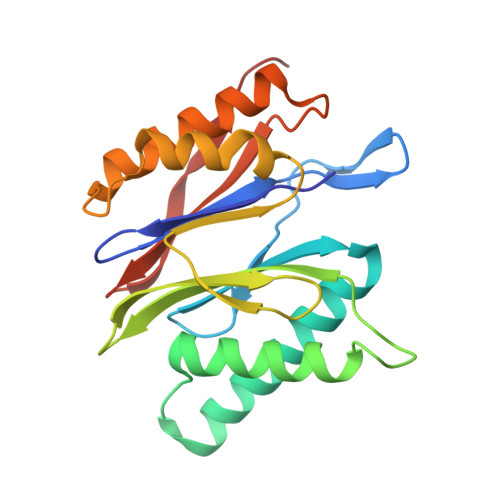
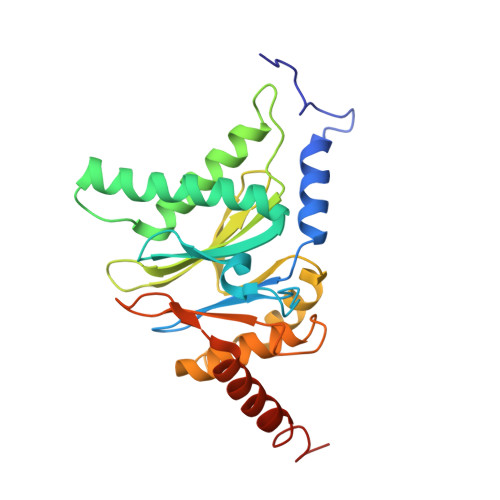
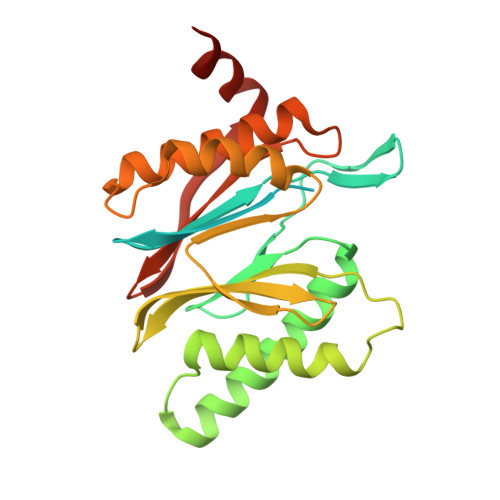
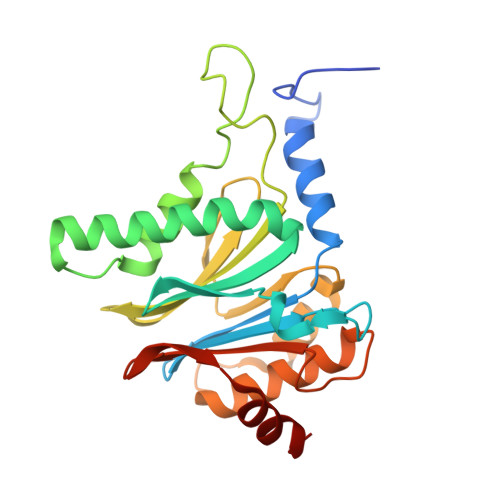
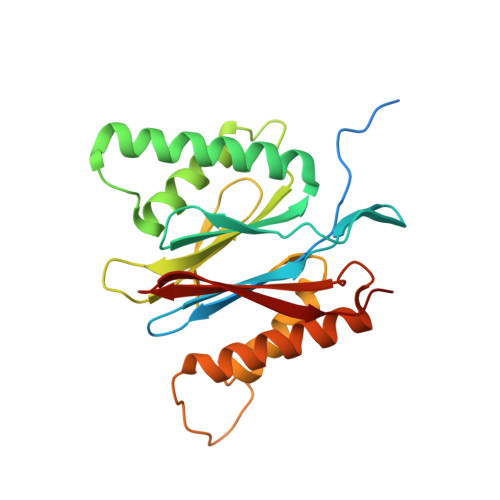
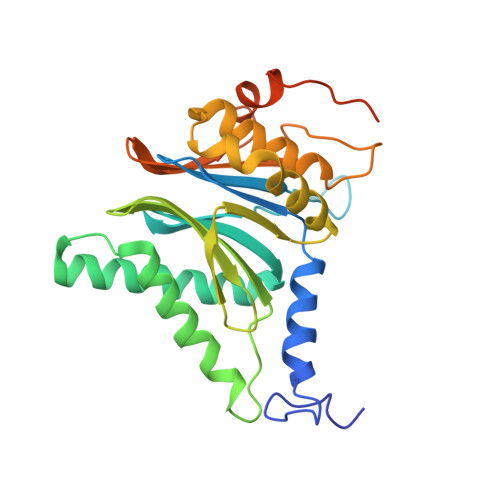
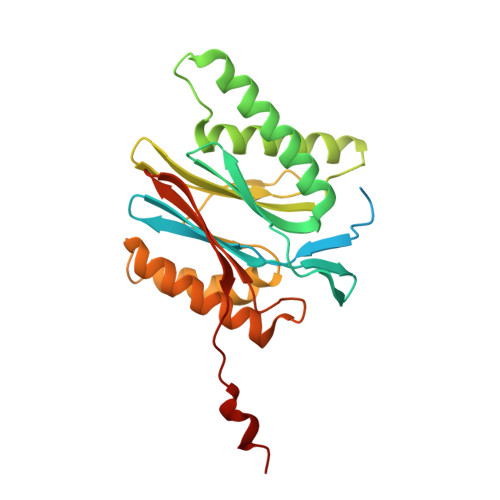
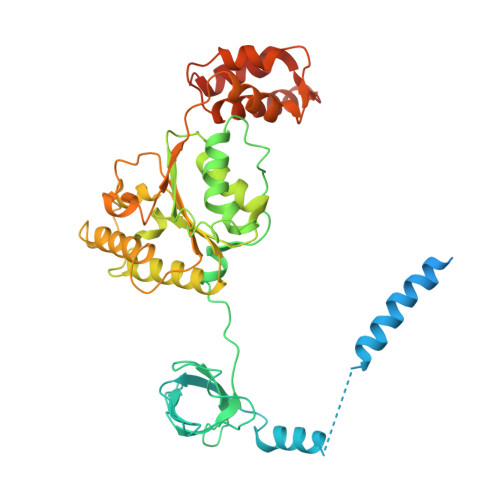
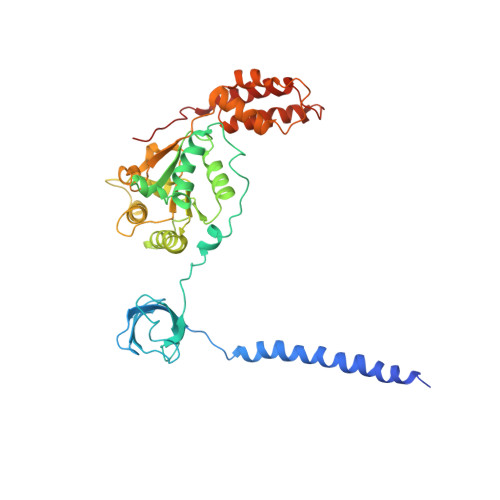
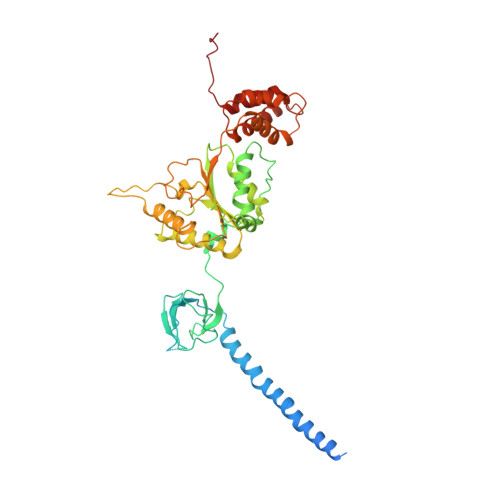
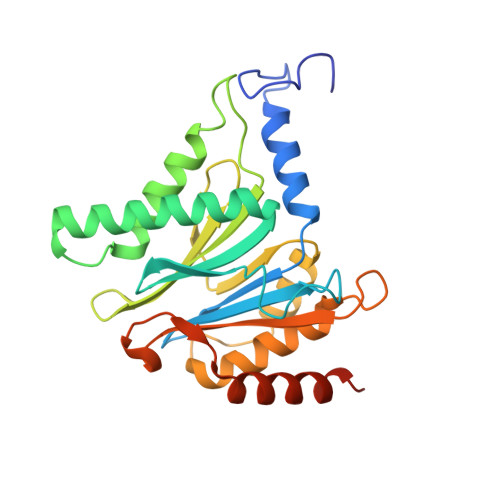
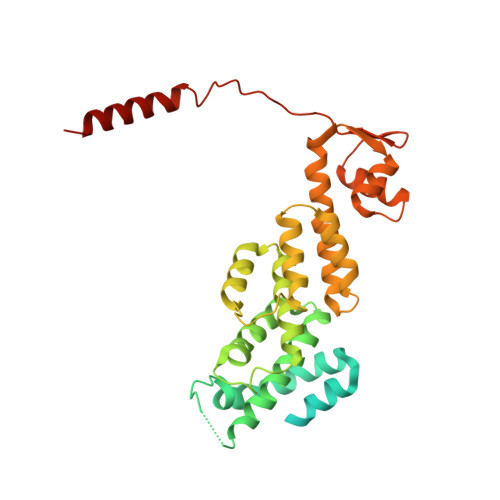
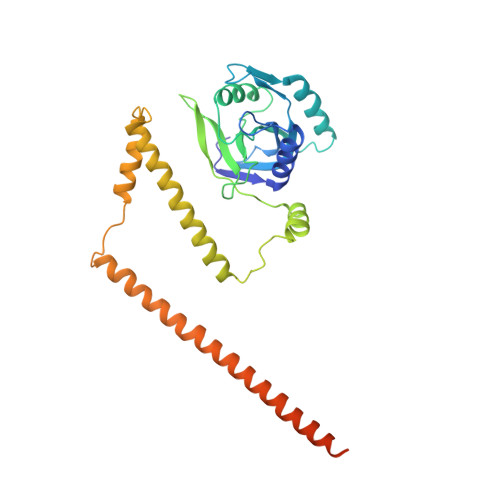
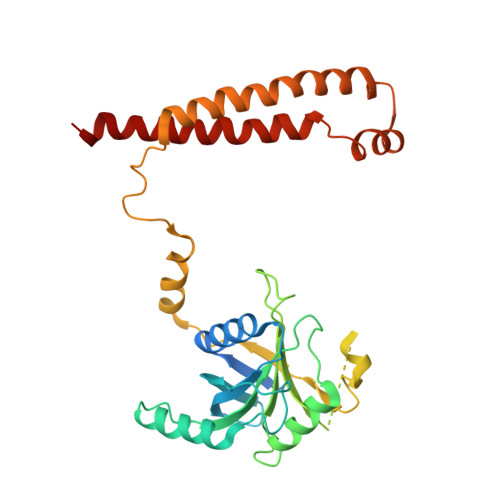
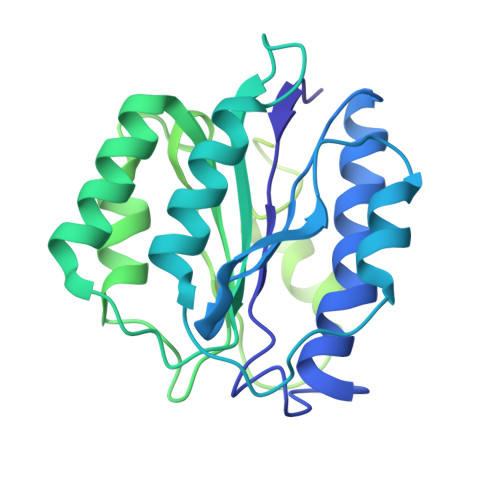
An atomic structure of the human 26S proteasome.
Huang, X., Luan, B., Wu, J., Shi, Y.(2016) Nat Struct Mol Biol 23: 778-785
- PubMed: 27428775 Search on PubMed
- DOI: https://doi.org/10.1038/nsmb.3273
- Primary Citation Related Structures:
5GJQ, 5GJR - PubMed Abstract:
We report the cryo-EM structure of the human 26S proteasome at an average resolution of 3.5 Å, allowing atomic modeling of 28 subunits in the core particle (CP) and 18 subunits in the regulatory particle (RP). The C-terminal residues of Rpt3 and Rpt5 subunits in the RP can be seen inserted into surface pockets formed between adjacent α subunits in the CP. Each of the six Rpt subunits contains a bound nucleotide, and the central gate of the CP α-ring is closed despite RP association. The six pore 1 loops in the Rpt ring are arranged similarly to a spiral staircase along the axial channel of substrate transport, which is constricted by the pore 2 loops. We also determined the cryo-EM structure of the human proteasome bound to the deubiquitinating enzyme USP14 at 4.35-Å resolution. Together, our structures provide a framework for mechanistic understanding of eukaryotic proteasome function.
- Ministry of Education Key Laboratory of Protein Science, School of Life Sciences, Tsinghua University, Beijing, China.
Organizational Affiliation: