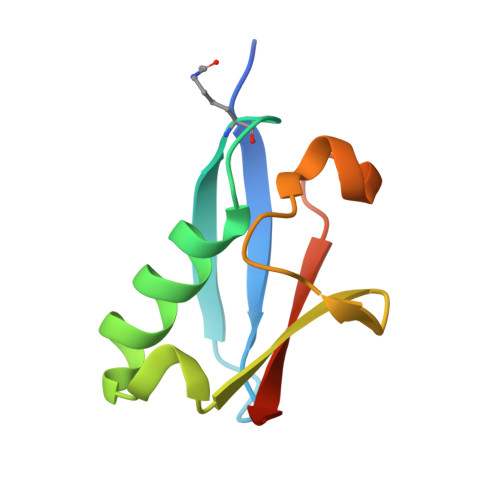
Acetylation of SUMO1 Alters Interactions with the SIMs of PML and Daxx in a Protein-Specific Manner.
Mascle, X.H., Gagnon, C., Wahba, H.M., Lussier-Price, M., Cappadocia, L., Sakaguchi, K., Omichinski, J.G.(2020) Structure 28: 157-168.e5
- PubMed: 31879127 Search on PubMed
- DOI: https://doi.org/10.1016/j.str.2019.11.019
- Primary Citation Related Structures:
6UYO, 6UYP, 6UYQ, 6UYR, 6UYS, 6UYT, 6UYU, 6UYV, 6UYX, 6UYY, 6UYZ - PubMed Abstract:
The interactions between SUMO proteins and SUMO-interacting motif (SIM) in nuclear bodies formed by the promyelocytic leukemia (PML) protein (PML-NBs) have been shown to be modulated by either phosphorylation of the SIMs or acetylation of SUMO proteins. However, little is known about how this occurs at the atomic level. In this work, we examined the role that acetylation of SUMO1 plays on its binding to the phosphorylated SIMs (phosphoSIMs) of PML and Daxx. Our results demonstrate that SUMO1 binding to the phosphoSIM of either PML or Daxx is dramatically reduced by acetylation at either K39 or K46. However, acetylation at K37 only impacts binding to Daxx. Structures of acetylated SUMO1 variants bound to the phosphoSIMs of PML and Daxx demonstrate that there is structural plasticity in SUMO-SIM interactions. The plasticity observed in these structures provides a robust mechanism for regulating SUMO-SIM interactions in PML-NBs using signaling generated post-translational modifications.
- Département de Biochimie et Médicine Moléculaire, Université de Montréal, C.P. 6128 Succursale Centre-Ville, Montréal, QC H3C 3J7, Canada.
Organizational Affiliation: