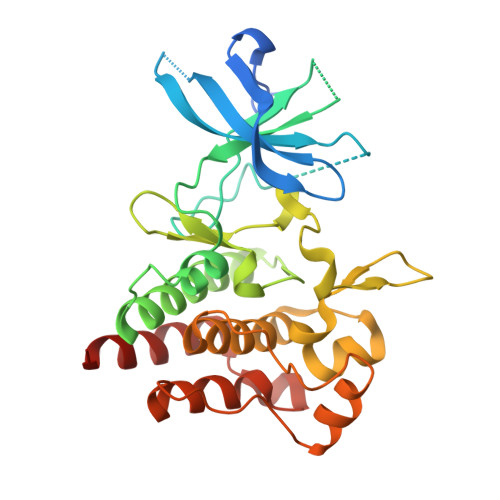
Structural Basis for Targeting the Folded P-Loop Conformation of c-MET.
Collie, G.W., Michaelides, I.N., Embrey, K., Stubbs, C.J., Borjesson, U., Dale, I.L., Snijder, A., Barlind, L., Song, K., Khurana, P., Phillips, C., Storer, R.I.(2021) ACS Med Chem Lett 12: 162-167
- PubMed: 33488978 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1021/acsmedchemlett.0c00392
- Primary Citation Related Structures:
7B3Q, 7B3T, 7B3V, 7B3W, 7B3Z, 7B40, 7B41, 7B42, 7B43, 7B44 - PubMed Abstract:
We report here a fragment screen directed toward the c-MET kinase from which we discovered a series of inhibitors able to bind to a rare conformation of the protein in which the P-loop adopts a collapsed, or folded, arrangement. Preliminary SAR exploration led to an inhibitor ( 7 ) with nanomolar biochemical activity against c-MET and promising cell activity and kinase selectivity. These findings increase our structural understanding of the folded P-loop conformation of c-MET and provide a sound structural and chemical basis for further investigation of this underexplored yet potentially therapeutically exploitable conformational state.
- Discovery Sciences, R&D, AstraZeneca, Cambridge, United Kingdom.
Organizational Affiliation: