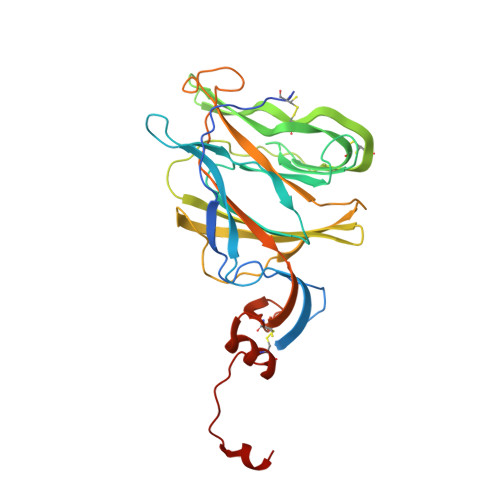
SARS-CoV-2 can recruit a heme metabolite to evade antibody immunity.
Rosa, A., Pye, V.E., Graham, C., Muir, L., Seow, J., Ng, K.W., Cook, N.J., Rees-Spear, C., Parker, E., Dos Santos, M.S., Rosadas, C., Susana, A., Rhys, H., Nans, A., Masino, L., Roustan, C., Christodoulou, E., Ulferts, R., Wrobel, A.G., Short, C.E., Fertleman, M., Sanders, R.W., Heaney, J., Spyer, M., Kjaer, S., Riddell, A., Malim, M.H., Beale, R., MacRae, J.I., Taylor, G.P., Nastouli, E., van Gils, M.J., Rosenthal, P.B., Pizzato, M., McClure, M.O., Tedder, R.S., Kassiotis, G., McCoy, L.E., Doores, K.J., Cherepanov, P.(2021) Sci Adv 7
- PubMed: 33888467 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1126/sciadv.abg7607
- Primary Citation Related Structures:
7B62, 7NT9, 7NTA, 7NTC - PubMed Abstract:
The coronaviral spike is the dominant viral antigen and the target of neutralizing antibodies. We show that SARS-CoV-2 spike binds biliverdin and bilirubin, the tetrapyrrole products of heme metabolism, with nanomolar affinity. Using cryo-electron microscopy and x-ray crystallography, we mapped the tetrapyrrole interaction pocket to a deep cleft on the spike N-terminal domain (NTD). At physiological concentrations, biliverdin significantly dampened the reactivity of SARS-CoV-2 spike with immune sera and inhibited a subset of neutralizing antibodies. Access to the tetrapyrrole-sensitive epitope is gated by a flexible loop on the distal face of the NTD. Accompanied by profound conformational changes in the NTD, antibody binding requires relocation of the gating loop, which folds into the cleft vacated by the metabolite. Our results indicate that SARS-CoV-2 spike NTD harbors a dominant epitope, access to which can be controlled by an allosteric mechanism that is regulated through recruitment of a metabolite.
- Chromatin Structure and Mobile DNA Laboratory, The Francis Crick Institute, London, UK.
Organizational Affiliation: