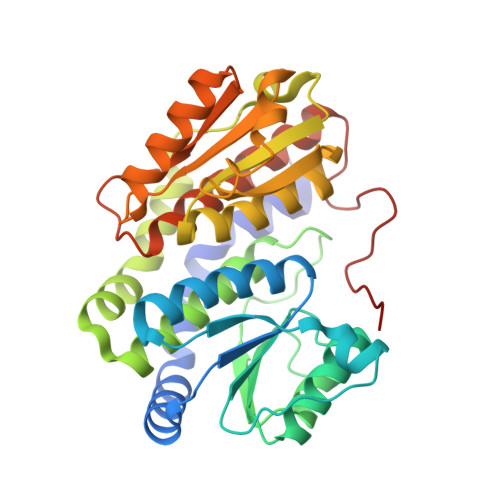
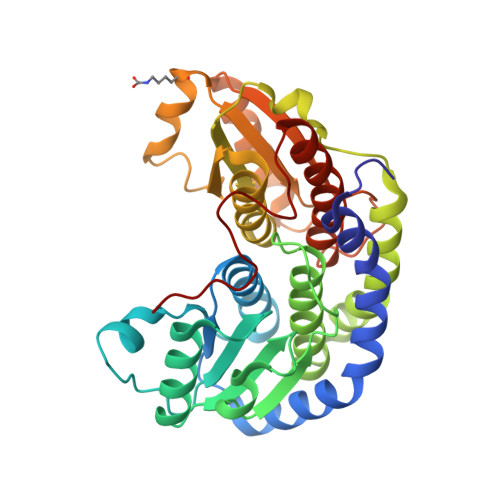
Substrate binding in the allosteric site mimics homotropic cooperativity in the SIS-fold glucosamine-6-phosphate deaminases.
Marcos-Viquez, J., Rodriguez-Hernandez, A., Alvarez-Anorve, L.I., Medina-Garcia, A., Plumbridge, J., Calcagno, M.L., Rodriguez-Romero, A., Bustos-Jaimes, I.(2023) Protein Sci 32: e4651-e4651
- PubMed: 37145875 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1002/pro.4651
- Primary Citation Related Structures:
8EOL, 8EYM, 8FDB - PubMed Abstract:
Glucosamine-6-phosphate (GlcN6P) deaminases from Escherichia coli (EcNagBI) and Shewanella denitrificans (SdNagBII) are special examples of what constitute nonhomologous isofunctional enzymes due to their convergence, not only in catalysis, but also in cooperativity and allosteric properties. Additionally, we found that the sigmoidal kinetics of SdNagBII cannot be explained by the existing models of homotropic activation. This study describes the regulatory mechanism of SdNagBII using enzyme kinetics, isothermal titration calorimetry (ITC), and X-ray crystallography. ITC experiments revealed two different binding sites with distinctive thermodynamic signatures: a single binding site per monomer for the allosteric activator N-acetylglucosamine 6-phosphate (GlcNAc6P) and two binding sites per monomer for the transition-state analog 2-amino-2-deoxy-D-glucitol 6-phosphate (GlcNol6P). Crystallographic data demonstrated the existence of an unusual allosteric site that can bind both GlcNAc6P and GlcNol6P, implying that the homotropic activation of this enzyme arises from the occupation of the allosteric site by the substrate. In this work we describe the presence of this novel allosteric site in the SIS-fold deaminases, which is responsible for the homotropic and heterotropic activation of SdNagBII by GlcN6P and GlcNAc6P, respectively. This study unveils an original mechanism to generate a high degree of homotropic activation in SdNagBII, mimicking the allosteric and cooperative properties of hexameric EcNagBI but with a reduced number of subunits.
- Departamento de Bioquímica, Facultad de Medicina, Universidad Nacional Autónoma de México (UNAM), Mexico City, Mexico.
Organizational Affiliation: