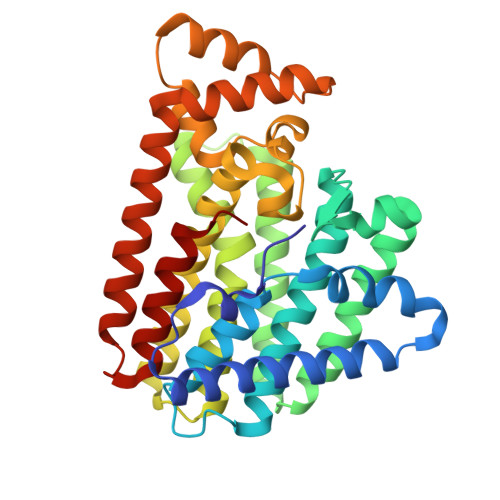
Structural insight of a bi-functional isoprenyl diphosphate synthase Rv0562 from Mycobacterium tuberculosis.
Wang, Q., Yang, Y., He, B., Huang, J.W., Kuo, C.J., Chen, C.C., Guo, R.T.(2025) Int J Biol Macromol 318: 145171-145171
- PubMed: 40513738 Search on PubMed
- DOI: https://doi.org/10.1016/j.ijbiomac.2025.145171
- Primary Citation Related Structures:
9IMQ, 9IMR, 9IMS, 9KUQ - PubMed Abstract:
Mycobacterium tuberculosis (Mtb) is the causative agent of tuberculosis. Mtb uses MK-9(II-H 2 ), which consists of an isoprenyl side chain containing nine isoprene units, with one being hydrogenated in the β-position, as an essential element in the electron transport system. Rv0562 that can operate geranylgeranyl diphosphate synthase (GGPPs) activity catalyzes the synthesis of MK-9(II-H 2 ) by condensing one molecule of DMAPP with eight molecules of isopentenyl diphosphate (IPP) to form a C 45 long-chain isoprenoid product. In this study, the structures of Rv0562 were determined in the apo-form at a resolution of 2.54 Å and in complex with IPP and Mg 2+ at a resolution of 1.89 Å, revealing detailed interactions between the enzyme and substrates. Moreover, the crystal structure of the Rv0562-DM variant was determined at 2.27 Å resolution in complex with polyethylene glycol (PEG), which occupies the substrate binding tunnel, mimicking the long-chain product. The chain length determination mechanism of Rv0562 is also probed through mutagenesis experiments. The obtained structures help us understand how Rv0562 catalyzes isoprenyl chain elongation, showing implications in developing new anti-Mtb treatments.
- State Key Laboratory of Biocatalysis and Enzyme Engineering, Hubei Hongshan Laboratory, School of Life Sciences, Hubei University, Wuhan 430062, PR China.
Organizational Affiliation: