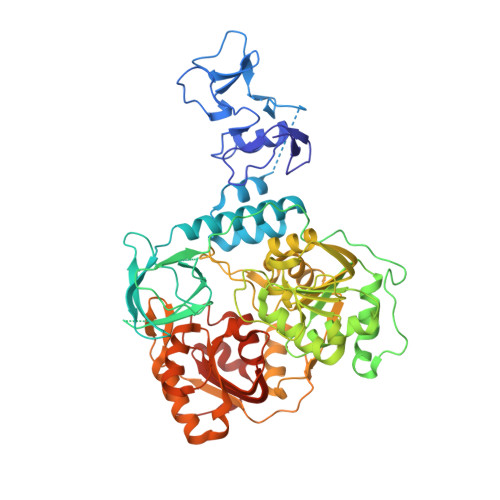
Structure, mechanism and crystallographic fragment screening of the SARS-CoV-2 NSP13 helicase.
Newman, J.A., Douangamath, A., Yadzani, S., Yosaatmadja, Y., Aimon, A., Brandao-Neto, J., Dunnett, L., Gorrie-Stone, T., Skyner, R., Fearon, D., Schapira, M., von Delft, F., Gileadi, O.(2021) Nat Commun 12: 4848-4848
- PubMed: 34381037 Search on PubMedSearch on PubMed Central
- DOI: https://doi.org/10.1038/s41467-021-25166-6
- Primary Citation Related Structures:
5RL6, 5RL7, 5RL8, 5RL9, 5RLB, 5RLC, 5RLD, 5RLE, 5RLF, 5RLG, 5RLH, 5RLI, 5RLJ, 5RLK, 5RLL, 5RLM, 5RLN, 5RLO, 5RLP, 5RLQ, 5RLR, 5RLS, 5RLT, 5RLU, 5RLV, 5RLW, 5RLY, 5RLZ, 5RM0, 5RM1, 5RM2, 5RM3, 5RM4, 5RM5, 5RM6, 5RM7, 5RM8, 5RM9, 5RMA, 5RMB, 5RMC, 5RMD, 5RME, 5RMF, 5RMG, 5RMH, 5RMI, 5RMJ, 5RMK, 5RML, ... Search all related entries - PubMed Abstract:
There is currently a lack of effective drugs to treat people infected with SARS-CoV-2, the cause of the global COVID-19 pandemic. The SARS-CoV-2 Non-structural protein 13 (NSP13) has been identified as a target for anti-virals due to its high sequence conservation and essential role in viral replication. Structural analysis reveals two "druggable" pockets on NSP13 that are among the most conserved sites in the entire SARS-CoV-2 proteome. Here we present crystal structures of SARS-CoV-2 NSP13 solved in the APO form and in the presence of both phosphate and a non-hydrolysable ATP analog. Comparisons of these structures reveal details of conformational changes that provide insights into the helicase mechanism and possible modes of inhibition. To identify starting points for drug development we have performed a crystallographic fragment screen against NSP13. The screen reveals 65 fragment hits across 52 datasets opening the way to structure guided development of novel antiviral agents.
- Centre for Medicines Discovery, University of Oxford, Oxford, UK. Joseph.Newman@cmd.ox.ac.uk.
Organizational Affiliation: